Sabitlenmiş Tweet

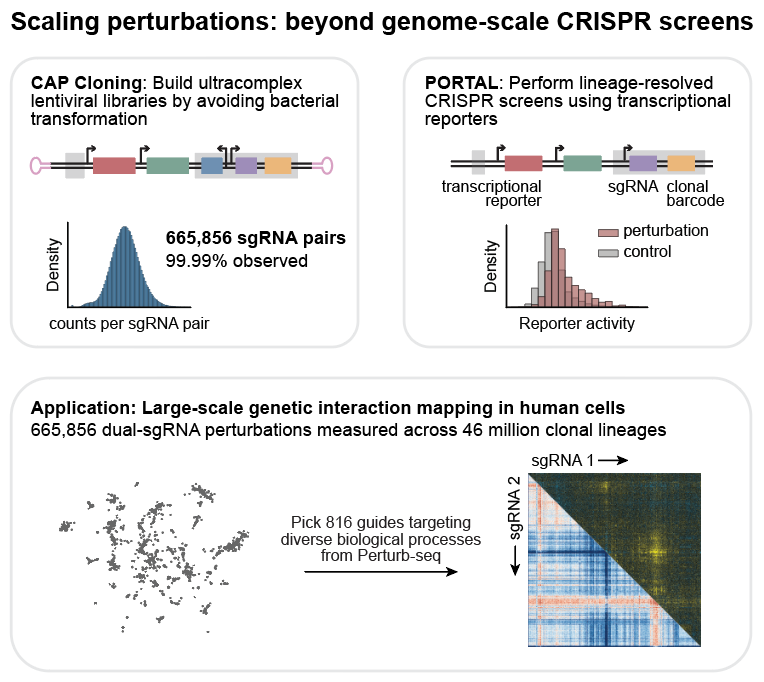
Happy to share that the paper on enhancer interactions led by @zrcjessica and I from my time in the @GrahamMcVicker lab is now published in Cell Genomics!
cell.com/cell-genomics/…
A thread on some of the new analyses below: (1/n)
English