
Biomni
55 posts


Biomni
@ProjectBiomni
The first IBE - collaborative agents designed for biologists to make more discoveries, faster, with rigor, and at scale. Built by @phylo_bio Stanford Spin-out



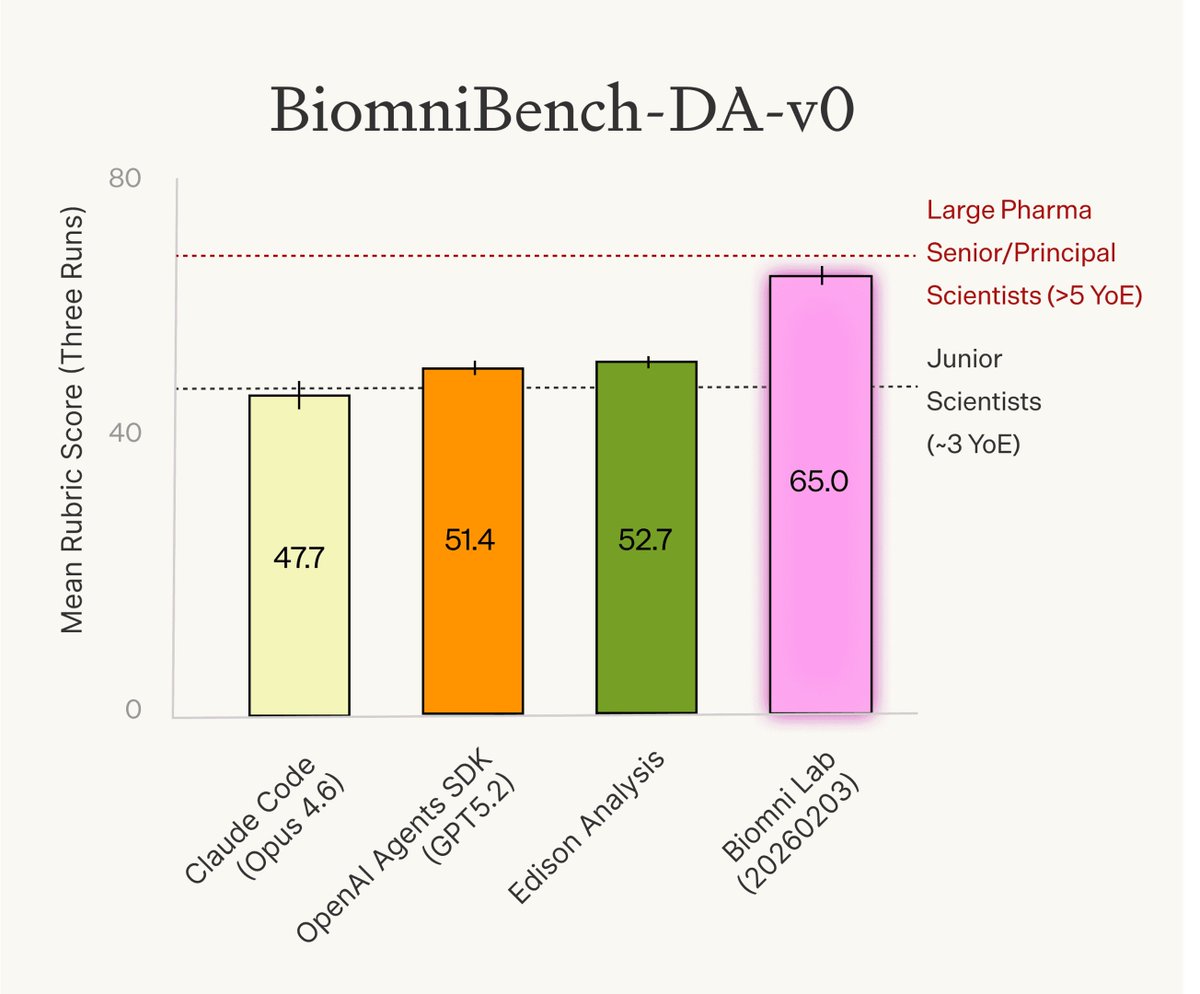
New research from Phylo: We rigorously evaluated today’s evals for biology agents and identified major issues: under-specified questions, incorrect ground truth, and — most importantly — evaluations that focus only on the final answer rather than the analytical process. To address this, we: - Refined an existing benchmark into BixBench-Verified-50 - Introduced BiomniBench, the first trace-based, real-world evaluation for AI agents in biology BiomniBench evaluates not just the output, but the full analytical workflow: data handling, method selection, statistical rigor, and biological interpretation. This mirrors how scientists evaluate each other’s work — and how agents should be evaluated too. Preliminary results: Biomni Lab achieves the strongest performance across both benchmarks. It performs on par with senior/principal scientists from large pharma companies (>5 years of experience) and surpasses junior scientists (~3 years of experience). We’ve open-sourced BixBench-Verified-50, and will release the full BiomniBench suite in the coming weeks. Read the full technical report: phylo.bio/blog/evaluatin…


Today we’re launching Phylo, a research lab studying agentic biology, backed by a $13.5M seed round co-led by @a16z and @MenloVentures / Anthology Fund @AnthropicAI. We’re also introducing a research preview of Biomni Lab, the first Integrated Biology Environment (IBE), where we’re imagining a new way biologists work. Biomni Lab uses agents to orchestrate hundreds of biological databases, software tools, molecular AI models, expert workflows, and even external research services in one workspace, supporting research end-to-end from question to experiment to result. Agents handle the mechanics, while you define the question, then review, steer, and decide. Scientists end up spending more time on science: asking questions, understanding mechanisms, and eliminating diseases. Phylo (@phylo_bio) is a spin-out of @ProjectBiomni, where we will maintain the open-source community and push open-science research. I’m grateful to continue building with my co-founders @YuanhaoQ @jure @lecong and the dream founding team @serena2z @TianweiShe @huangzixin20151 @gm2123 @margaretwhua @malayhgandhi. We’re also fortunate to be advised by leading scientists @zhangf, Carolyn Bertozzi, and @fabian_theis, and supported by an amazing group of investors including @JorgeCondeBio @zakdoric Matt Kraning @ZettaVentures @dreidco @conviction @saranormous @svangel @valkyrie_vc and others. Biomni Lab is available for free today: biomni.phylo.bio Learn more in our launch post: phylo.bio/blog/company-f… We are also hosting launch events - join us at South San Francisco: luma.com/n8k8qb0n Virtual: luma.com/l5ryjaij We’re also hiring! phylo.bio/careers

We are excited to partner with @Ginkgo Datapoints to demonstrate how Biomni can accelerate functional genomics analysis. Biomni Lab analyzed 10+ transcriptomic and Cell Painting datasets from the Ginkgo Datapoints (GDPx) collection in hours instead of weeks, with results validated by Ginkgo scientists. Try it today at biomni.phylo.bio and learn more at: phylo.bio/blog/ginkgo-bi… @jrkelly

Today we’re launching Phylo, a research lab studying agentic biology, backed by a $13.5M seed round co-led by @a16z and @MenloVentures / Anthology Fund @AnthropicAI. We’re also introducing a research preview of Biomni Lab, the first Integrated Biology Environment (IBE), where we’re imagining a new way biologists work. Biomni Lab uses agents to orchestrate hundreds of biological databases, software tools, molecular AI models, expert workflows, and even external research services in one workspace, supporting research end-to-end from question to experiment to result. Agents handle the mechanics, while you define the question, then review, steer, and decide. Scientists end up spending more time on science: asking questions, understanding mechanisms, and eliminating diseases. Phylo (@phylo_bio) is a spin-out of @ProjectBiomni, where we will maintain the open-source community and push open-science research. I’m grateful to continue building with my co-founders @YuanhaoQ @jure @lecong and the dream founding team @serena2z @TianweiShe @huangzixin20151 @gm2123 @margaretwhua @malayhgandhi. We’re also fortunate to be advised by leading scientists @zhangf, Carolyn Bertozzi, and @fabian_theis, and supported by an amazing group of investors including @JorgeCondeBio @zakdoric Matt Kraning @ZettaVentures @dreidco @conviction @saranormous @svangel @valkyrie_vc and others. Biomni Lab is available for free today: biomni.phylo.bio Learn more in our launch post: phylo.bio/blog/company-f… We are also hosting launch events - join us at South San Francisco: luma.com/n8k8qb0n Virtual: luma.com/l5ryjaij We’re also hiring! phylo.bio/careers


Today we’re launching Phylo, a research lab studying agentic biology, backed by a $13.5M seed round co-led by @a16z and @MenloVentures / Anthology Fund @AnthropicAI. We’re also introducing a research preview of Biomni Lab, the first Integrated Biology Environment (IBE), where we’re imagining a new way biologists work. Biomni Lab uses agents to orchestrate hundreds of biological databases, software tools, molecular AI models, expert workflows, and even external research services in one workspace, supporting research end-to-end from question to experiment to result. Agents handle the mechanics, while you define the question, then review, steer, and decide. Scientists end up spending more time on science: asking questions, understanding mechanisms, and eliminating diseases. Phylo (@phylo_bio) is a spin-out of @ProjectBiomni, where we will maintain the open-source community and push open-science research. I’m grateful to continue building with my co-founders @YuanhaoQ @jure @lecong and the dream founding team @serena2z @TianweiShe @huangzixin20151 @gm2123 @margaretwhua @malayhgandhi. We’re also fortunate to be advised by leading scientists @zhangf, Carolyn Bertozzi, and @fabian_theis, and supported by an amazing group of investors including @JorgeCondeBio @zakdoric Matt Kraning @ZettaVentures @dreidco @conviction @saranormous @svangel @valkyrie_vc and others. Biomni Lab is available for free today: biomni.phylo.bio Learn more in our launch post: phylo.bio/blog/company-f… We are also hosting launch events - join us at South San Francisco: luma.com/n8k8qb0n Virtual: luma.com/l5ryjaij We’re also hiring! phylo.bio/careers






Meet Slingshots // One. This inaugural batch includes leading-edge researchers advancing the science and practice of AI - with benchmarks, frameworks, and agents that ship real impact into the world. We're honored to support research from: @alexgshaw @Mike_A_Merrill @lschmidt3 @andykonwinski @lateinteraction @a1zhang @GregKamradt @mikeknoop @fchollet @jyangballin @KLieret @Diyi_Yang @tyler_griggs_ @profjoeyg @istoica05 @DachengLi177 @shiyi_c98 @matei_zaharia @sijun_tan @ralucaadapopa @ying11231 @lm_zheng @Tim_Dettmers @Muennighoff @AndrewYNg @YejinChoinka @atharva_sehgal @yisongyue @ChengleiSi @tatsu_hashimoto @KexinHuang5 @lisabdunlap @JacobSteinhardt @LakshyAAAgrawal This first batch sets a VERY high bar, and we’re proud to call these researchers part of the Laude community. More details here: laude.org/updates/slings…



From Code to Cure Webinar🧬 Join our own @ElmuratovArtem and @Stanford’s @lecong on Oct 23 at 9 AM PDT for a webinar exploring how AI agents are accelerating biomedical discovery from CRISPR-GPT to RNAGenesis. Sign up: nebius.com/events/webinar… #AI #Biotech #Webinar




