2438@ROM retweetledi

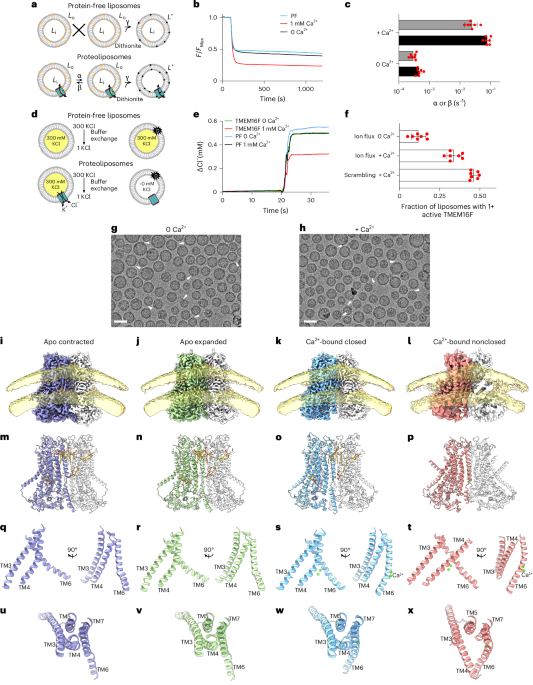
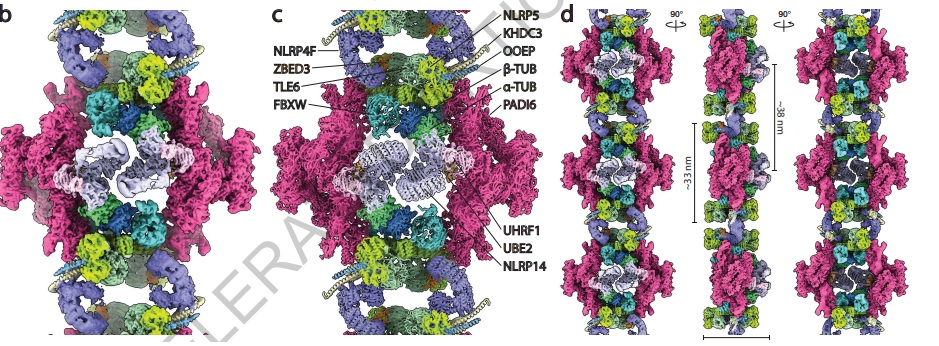
"Cytoplasmic lattices are megadalton storage complexes in mammalian oocytes."
Uh yeah, totally normal, no oocyte-specific alien spaceships here:
@Nature #oocyte #ScienceWithTinyThings
tinyurl.com/mra3dx6t
English
2438@ROM
54.5K posts






中国人と韓国人に聞きたいんだけど、他の国の箸は使いづらいとか思ってるの? 私は中国の箸は長くて慣れないし、韓国の鉄の箸も慣れない。 だから逆に日本の箸も他の箸文化の国からしたら使いづらいんじゃないかなって疑問なんだよね。 他文化をディスる意味ではなくただ意見を聞いてみたい。





自治体のIT機器、中国製品を排除 政府の認定品のみ使用可能に nikkei.com/article/DGXZQO…




台湾の主要政党 民進党…「台湾への愛」以外の部分はガバガバに雑で軽い 国民党…「民国への愛」以外の部分はガバガバに雑で軽い 民衆党…上記両党に嫌気が差した人に支持され、台湾への愛や民国への愛すらガバガバに雑で軽い

自治体のIT機器、中国製品を排除 政府の認定品のみ使用可能に nikkei.com/article/DGXZQO…