
It's #MatchDay! Please join us in welcoming our 2026-2027 intern class! Welcome to the Stanford Department of Medicine! @StanfordMedRes is thrilled to have you! ❤️🥳🎉🤩🍾 @Ron_Witteles @StanfordDeptMed #Match2026
Zachary Flamholz
139 posts

@zflam94
current student, future physician scientist | bringing data to medicine | opinions are my own | RT ≠ endorsement

It's #MatchDay! Please join us in welcoming our 2026-2027 intern class! Welcome to the Stanford Department of Medicine! @StanfordMedRes is thrilled to have you! ❤️🥳🎉🤩🍾 @Ron_Witteles @StanfordDeptMed #Match2026



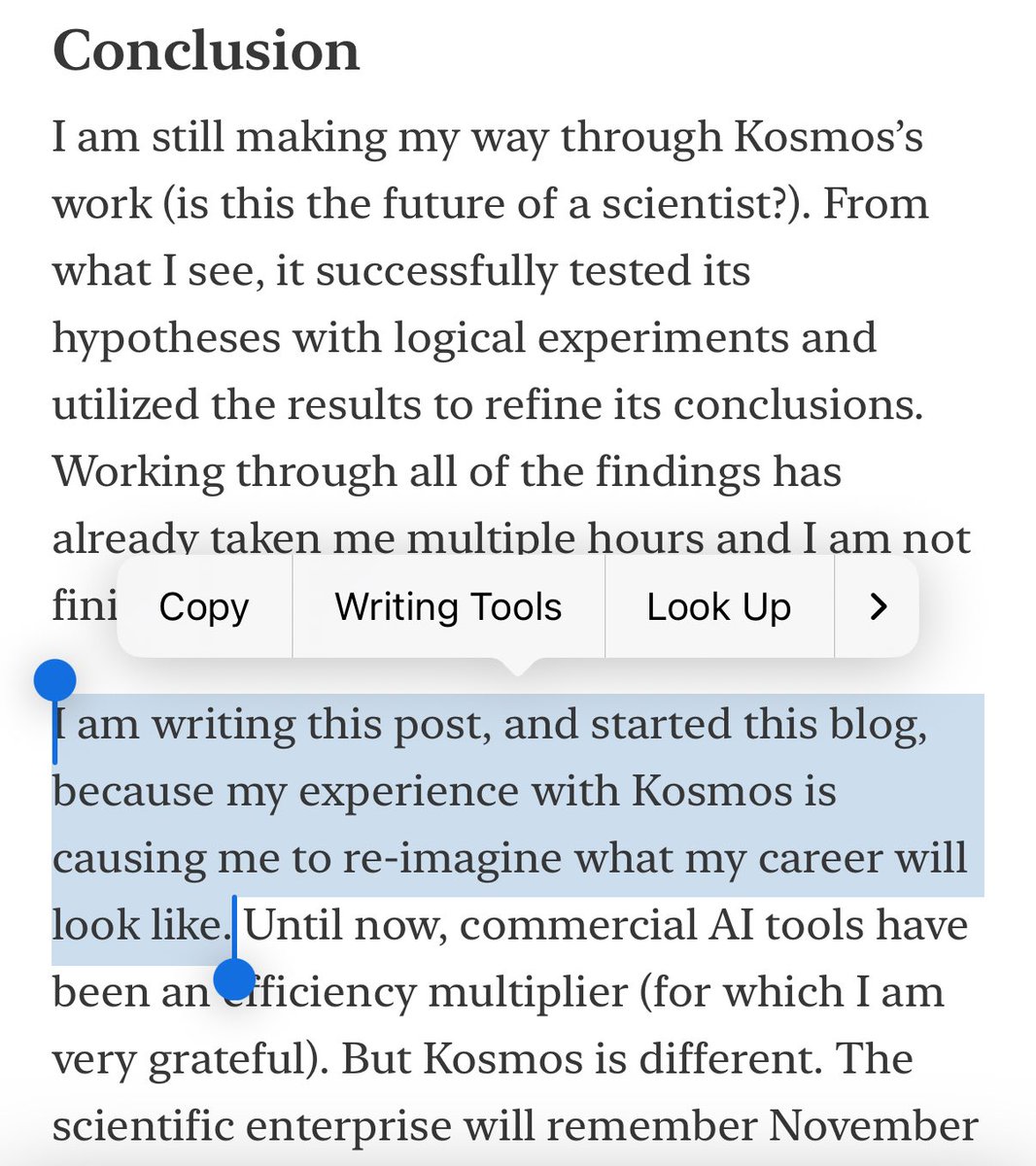
Science is too slow. At Edison, we are integrating AI Scientists into the full stack of research, from basic discovery to clinical trials. We want cures for all diseases by mid-century. We have raised a $70M seed to get started. Join us. We need cracked software engineers who want to work on finding cures rather than selling ads and generating slop. If you’re reading this, you’re probably a candidate. We need brilliant AI researchers who want to figure out how AI will accelerate real-world science. We need scientists and researchers with deep expertise in biology, biotech, and pharma who want to figure out how to integrate AI deeply into scientific workflows, from ideation to experimentation, and how to measure success or failure. We need extraordinarily talented generalist operators across BD, sales, product management, and partnerships who can focus on getting our tools into the hands of pharmaceutical companies. If any of these roles sound like you, get in touch. We are also expanding access to our platform. Our goal is to accelerate science writ large. To that end, we will continue to give academics and students 650 credits/mo indefinitely. I can’t promise we’ll keep this up forever, but we will try. Kosmos will still cost 200 credits, and the other agents (Analysis, Literature, etc.) will cost 1 or 2 credits. All paid users will have access to our regular agents, like our Analysis agent, Literature agent, and so on, for free via the UI. API access will still be paid, and users without a paid subscription will continue to get 10 credits per month for those agents. Our $200/mo subscription for 650 credits/mo is staying in place for now, but might be phased out at our next major product update. Along the lines of accelerating science, we’re also doing a major release of PaperQA today, our flagship open source literature agent, as part of our commitment to open science. In the short run, expect major improvements to Kosmos, including the ability to automatically access data, the ability to steer its exploration, and the ability to converse directly with its world model. In the long run, expect exponentially increasing rates of scientific discoveries, in biology and elsewhere. Our round is led by Triatomic Capital, Spark Capital, and a major US institutional biotech investor. We are also joined in this round by existing investors Pillar VC and Susa Ventures, two exceptional early-stage funds who backed us at founding, along with Striker Venture Partners, Hawktail VC, Olive VC, and a host of exceptional angels that includes famous AI researchers, the CEOs of multiple frontier AI labs, and leadership of major biotech and pharma companies.











Interested in phages? Live in the Bay Area & feel like spending a morning at Stanford? You're invited to a special phage seminar with 2 of my favorite phage profs, visiting us all the way from Europe: Speakers: Joana Azeredo (University of Minho) @azeredo_joana & Martin J. Loessner (ETH Zurich) @MartinLoessner Come learn about engineered phages, phage therapy, biofilms, antibiotic resistance, & more. When & Where: Jan 31, 2025, from 11am-12pm at Munzer Auditorium at Stanford Hope to see you there!


Whoa!! @EvoscaleAI just came out with a potentially useful upgrade to our favorite ESM-2 pLM with three new ESM C representation models: 300M, 600M, and 6B! 😳 Here, I'll do three semi-analyses based on the release: model modifications, "evaluations", and license. Would be happy if the ESM team corrects me on any misunderstandings! 😅 Model Modifications (😍) -Context length of 2048!! We can represent bigger Cas9s and other multi-domain proteins now! ✂️🧬 -Larger training set than the filtered ~65 million for ESM-2, especially with a lot of new metagenomic sequences! UniRef + MGnify + JGI data -- beautiful! 🦠 -Pre-Attention LayerNorm for query-key matrices -- should be good for improving attention stability. 🏂 -SwiGLU will def improve parameter efficiency over GELU in the FFN layers! 🦾 -RoPE should give improved performance on longer, more structurally complex proteins. 😉 -I'm just glad ESM-C does not require structural input -- gosh, I just REALLY disliked that about ESM3 and all of the new structure-infused pLMs. 😅 -Overall, this is a simple, sequence-based transformer with strong modifications. Love the work! 👏👏 Evaluations (😒) -Okay, guys, really? Emergent structure -- that's the benchmark you open with? 🙄 -Yes, emergent structure is nice because this is sequence-only and has no structural supervision or structurally-biased latents. 🙅♂️ And the scaling laws do look good! -BUT what about variant effects, solubility, fluorescence, localization, functional properties, binding affinity? 😔 All the reasons people love ESM! -Fingers crossed, we'll see these in the preprint to compare against ESM-2 🤞 (unlike the ESM3 benchmarks), otherwise we'll be having to do these benchmarks for the 300M version against ESM-2-650M. 😓 -Speaking of the 300M version... License (😬😬😬) -I did a deeper look, and yes, 300M is "fully" open (both to academics and industry folks)! If it is comparably strong to ESM-2-650M, this is good news. 🌟 -600M and of course, 6B, are off-limits for academic labs like mine who develop therapeutics with pLMs and may try bring to market later. 🙅♂️ Non-commercial use only so be careful! 😬 -Back to 300M, if you DO use it, you need to attribute it as "Generated with ESM," include "Built with ESM" in public-facing outputs (have ESM in the model name), and credit ESM in regulatory or public disclosures. ✏️ Otherwise, you break the licensing agreement. ☠️ My verdict: -Look guys, ESM-2 is completely free. Unless ESM C-300M does substantially better on function/variant prediction and other relevant benchmarks, it's probably not worth changing and dealing with the license terms. 🫤 -My lab will likely stick (not set in stone) to creating relevant and useful variants of ESM-2-650M (or even @pengzhangzhi1's new FlashAttention-based ESM-2-650M: github.com/pengzhangzhi/f…), which is a perfectly potent and efficient starting point for most downstream prediction and design tasks. Why fix what's not broken? 🤷♂️ By the way, as some updates, we're getting closer to publishing peer-reviewed, improved ESM-2 based models for: -Incorporating PTMs with PTM-Mamba: biorxiv.org/content/10.110… -Representing fusion oncoproteins with FusOn-pLM: biorxiv.org/cgi/content/sh… -Designing de novo target-binding peptides with PepPrCLIP (biorxiv.org/content/10.110…), PepMLM (arxiv.org/abs/2310.03842) -Designing motif-specific binders with moPPIt (biorxiv.org/content/10.110…) -Generating antimicrobial peptides with AMP-Diffusion (biorxiv.org/content/10.110…) -Designing membrane proteins with MeMDLM: arxiv.org/abs/2410.16735 -Designing metal binding proteins with MetaLATTE: biorxiv.org/content/10.110… AND SO MUCH MORE TO COME SOON! 🌟 So keep a lookout 👓 and thanks for following our work! 🥹


A warm welcome to Rockefeller, Avi! Avi Flamholz's lab will focus on the role of soil microbes in the carbon cycle, with the goal of unlocking new ways to harness the power of microbes to slow down or reverse rising global temperatures. rockefeller.edu/news/36919-new…


This is a foundational achievement. MMseqs2-GPU virtually *erases* the bottleneck of homology search in many workflows, including structure prediction. @sacdallago @milot_mirdita @thesteinegger, Felix, Bertil, and Alejandro too! developer.nvidia.com/blog/boost-alp…


