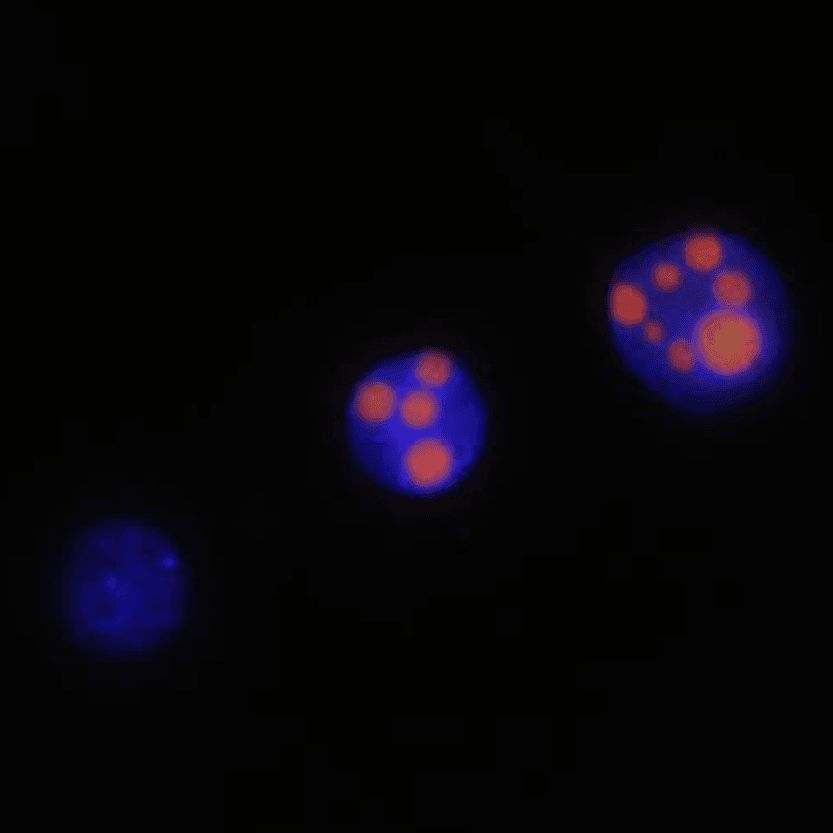
Ever wondered about the function of nucleoli in zygotes and early embryos, and their interaction with the embryonic genome? Mark your calendars! #Embryology #Genomics

English
Fulka Lab
115 posts


@FulkaLab
We are the Department of Cell nucleus plasticity, a part of Institute of Experimental Medicine of Czech academy of Sciencies. #oocyte #nucleolus #chromatin





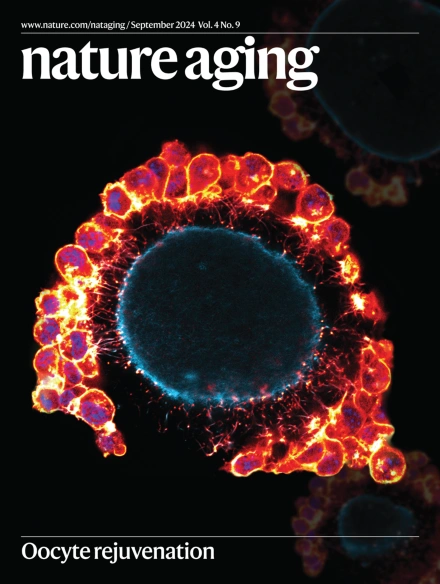
🗞️Our September issue is out!🎉 Read about cognitive decline, reproductive aging, mouse lifespan-expanding botanical extracts, ICOPE and much more🎨image credit: Hai Yang Wang nature.com/nataging/volum… pic.twitter.com/56dmXSg7tp