InnovativeBioresearch🇮🇹
6.7K posts


InnovativeBioresearch🇮🇹
@InnBioresearch
INNBC Chain | High-speed L1 Blockchain 🧬 Pioneering #DeSci & Gaming. Official upgrade of the legacy $INNBC 2018 asset #L1 #Web3


Ogni tanto #Steam riesce ancora a ricordare una cosa semplice: esistono giochi gratis che puoi scaricare e tenere per sempre, senza condizioni. spaziogames.it/notizie/8-picc…





People will be like “why is aspirin good for you” and I’m like.. how much time do you have
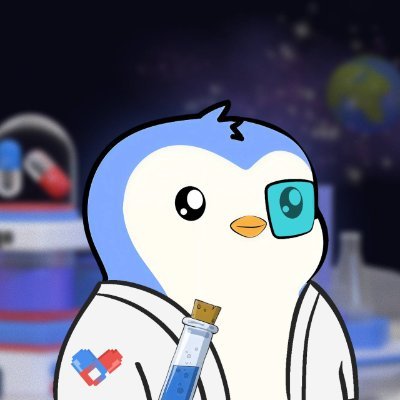





























This is a masterclass in 'bio/acc' PR, but let’s look at the actual science. Designing a molecule on a Sunday afternoon is not the same as discovering a drug. Here is why this is speculative marketing, not validated science: 1. Confusion between "Design" and "Discovery" Designing a molecule in silico (on a computer) that theoretically binds to a receptor like GLP-1 is relatively simple today thanks to tools like AlphaFold or molecular docking. • The Reality: 99% of molecules that look perfect on a screen fail the moment they touch a living cell. "Discovery" is not the digital structure; it is the demonstration of biological safety and efficacy, which cannot be done in an afternoon. 2. The Pharmacokinetics Wall (ADMET) The report mentions ADMET data (Absorption, Distribution, Metabolism, Excretion, and Toxicity) as if they were settled. • The Reality: AI predictions on toxicity are notoriously imprecise. The report itself admits a "DILI flag" (Drug-Induced Liver Injury) for candidate RLN-4522. In pharmacology, a liver toxicity signal usually kills a project immediately. Dismissing it as a "highlighted risk" in a post is a way to mask a technical failure as "scientific transparency." 3. The Cost Fallacy: $15,000 vs $50M Comparing a $15,000 "go/no-go" experiment to the $50M spent by Big Pharma is an "apples-to-oranges" fallacy. • The Reality: The $50M budget covers complex chemical synthesis, animal models (essential for metabolic drugs), formulation stability, and regulatory pre-clinical tests. Conducting a "metabolite trapping" test in liver microsomes is a standard, low-cost initial screen. It’s like claiming you built a skyscraper because you bought the bricks and made a 3D rendering. 4. Absence of Peer-Review Data❌❌❌ Validated science requires peer-review and the publication of raw data from physical laboratory (wet-lab) experiments. • The document shown is a "Pre-IND Computational Design"—essentially an AI-generated report analyzing its own output. It is a closed feedback loop. Without testing on complex organisms, one cannot claim a molecule will solve "gastroparesis" or "immunological risks" of existing drugs. 5. Conflict of Interest and DeSci (Decentralized Science) The author mentions @BioProtocol and @Molecule_sci, which are platforms looking for investments or issuing tokens. • The Goal: These posts create hype to inflate the perceived value of tech infrastructures or tokens, attracting retail investors who may lack the technical expertise to distinguish between a digital protein model and an actual medicine. Summary: This is PR because it shifts focus from the biological result (a drug that heals) to the technological process (an AI that generates reports)
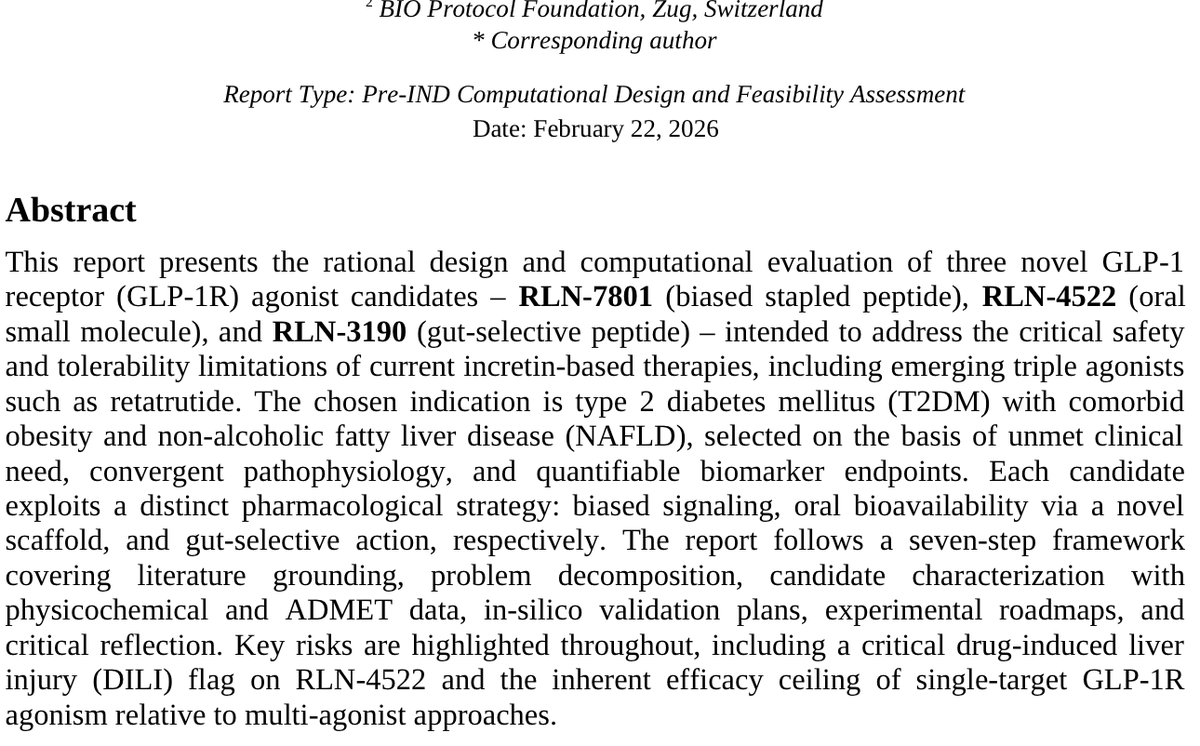




🧵 We just designed three novel GLP-1 receptor agonists from scratch on a Sunday afternoon using autonomous AI agents on @bio_protocol infrastructure Not a pharma lab. Not a $70M seed round. An AI scientist, a hypothesis on @ScienceBeach, and onchain compute via @Molecule_sci - here's what happened