Julian Naderi
25 posts

Julian Naderi
@Naderij_
Transcriptional Regulation & Protein engineering | PhD student @HniszLab @MPI_Molgen


Online now: The regulatory landscape of 5′ UTRs in translational control during zebrafish embryogenesis dlvr.it/THN8wk



Genetic evidence for functional specificity in biomolecular condensates: nature.com/articles/s4157… Free read only copy: rdcu.be/dXDvY @bsabari @idpsig @HymanLab @PappulabWashU @Pha_Tran_Papers @NatureRevGenet @MPI_MolGen #Condensates #IDR





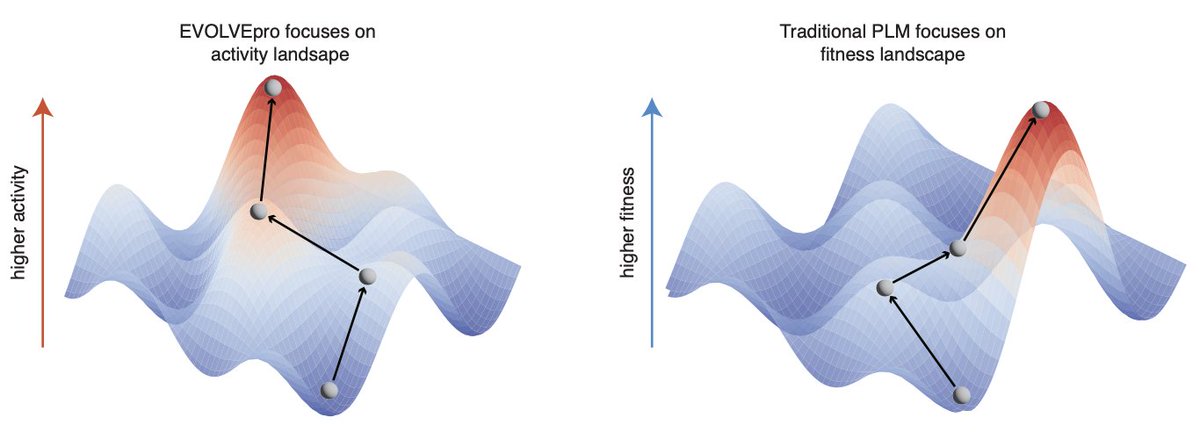
Directed evolution is key for unlocking new protein function But is difficult and time consuming So how can we accelerate protein design by 10-100x? With AI! Now introducing EVOLVEpro, an LLM-based model for evolving proteins rapidly and efficiently biorxiv.org/content/10.110…







