SheqLab
2.9K posts

SheqLab
@ShechnerLab
The Shechner lab in UW Pharmacology. We study Noncoding RNAs and cellular architecture. He/His/Him. All comments are my own. @ShechnerLab.bsky.social

Today we report single-cell APEX-seq (scAPEX-seq) — a new method for unbiased mapping of *subcellular* transcriptomes at single-cell resolution. This approach reveals cell states that are not detectable by standard scRNA-seq, and enabled us to identify regulators of CAR T function that improve solid tumor killing. biorxiv.org/cgi/content/sh…
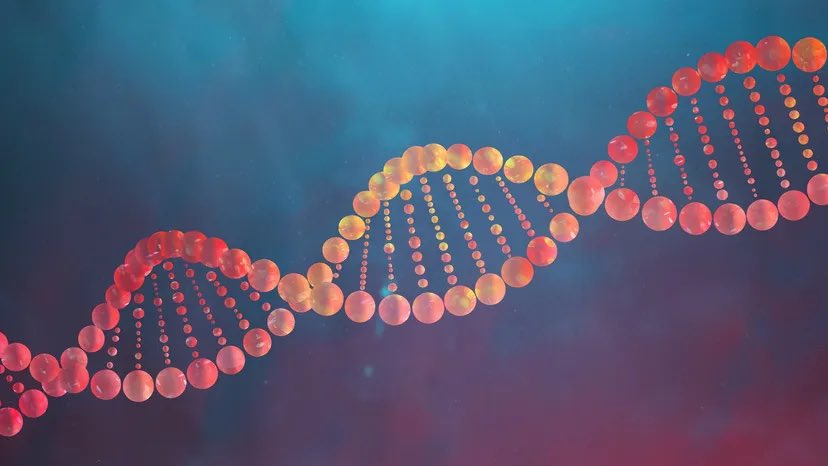


As James Watson, co - discoverer of the DNA double helix structure, has passed today (RIP), I want to address an important issue: most modern visualizations of the DNA double helix are actually WRONG. The original Watson-Crick "bands wrapped around a rod" picture depiction is the geometrically correct way to represent canonical B-DNA, whereas modern depictions usually look like the whole object is just a twisted ribbon. 🧬 (including this emoji). I only realized this myself a couple of years ago myself , when I read their original paper for the first time.

I’m thrilled to share our preprint that uses deep learning to interrogate structure-function relationships in condensates! biorxiv.org/content/10.110… In here, we ask: can AI read condensate biology from pictures alone? Turns out yes... see what we discover below! 🤖🧬🖼️ (1/10)





I'm thrilled to announce my next career step-- I’m joining Genentech as a Principal Scientist & Lab Head in Discovery Oncology! I’ll be hunting new ways to target cancers using my background in disordered nuclear proteins.

Please join us in congratulating @HsiehLab of @FredHutch for this 2025 Bladder Cancer Research Innovation Award. Dr. Hsieh’s research aims to uncover how mutations in other commonly altered genes contribute to diversity in BC. Click here to learn more: bcan.org/bcan-awards-60…

More than 2700 3′UTRs are highly conserved. These 3′UTRs are essential components in mRNA templates, as their deletion decreases protein activity without changing protein abundance. Highly conserved 3′UTRs help the folding of proteins with long IDRs. biorxiv.org/content/10.110…

New from the Rouhanifard Lab! What if we could sort single cells based on whether a specific gene was actively being transcribed—and then ask what drives that burst? Introducing: NuclampFISH










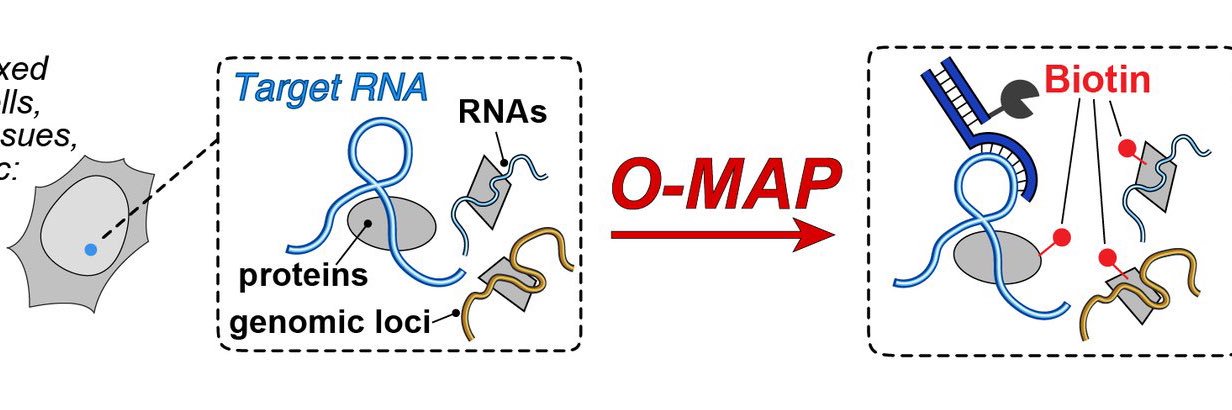
#RNAsequencing helps map #RNA interactions with proteins and #DNA, revealing cellular processes and disease mechanisms. The O-MAP method by the @ShechnerLab enables precise RNA neighborhood mapping for deeper insights. - @UW rna-seqblog.com/o-mapping-the-…



Our work on cellular RNA interactions with MAVS that promote antiviral signaling is finally live. So grateful to our indispensable collaborators as well as reviewers who helped to strengthen our work. #RNA #innateimmunity #virology @RamLabUW science.org/doi/10.1126/sc…