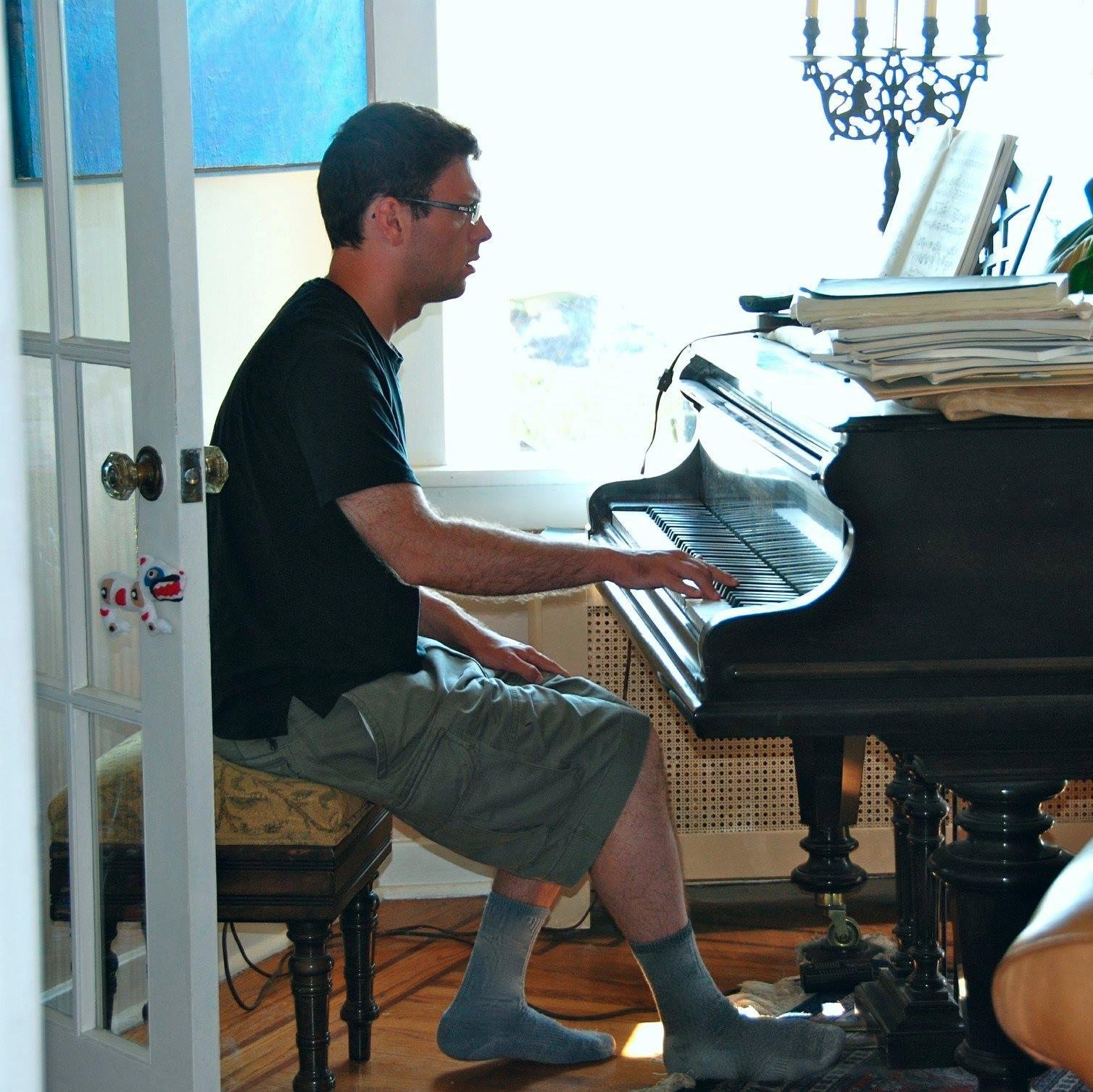
پن کیا گیا ٹویٹ
tbepler
131 posts

tbepler
@tbepler1
Scientist and Group Leader of the Simons Machine Learning Center @SEMC_NYSBC. Co-founder and CEO of https://t.co/YcCw0bpHZo. Opinions are my own.
شامل ہوئے Ekim 2018
152 فالونگ645 فالوورز
tbepler ری ٹویٹ کیا

We’re excited to announce our expanded partnership with Boehringer Ingelheim. Together, we are building the future of AI‑driven antibody discovery and optimization.
openprotein.ai/strategic-part…
English

@PatrickSoga @openprotein Yes, we will probably take 1-2 interns next year. If you're interested feel free to send us your CV
English

@tbepler1 @openprotein Are you taking any interns this spring/sunmer?
English

New job openings @openprotein across protein foundation model research, computational protein design, and cloud platform engineering
openprotein.ai/careers
English

@RippaSatss @openprotein We'll consider anyone with relevant skills who's motivated. Send us your CV and tell us more about what you've worked on!
English

Quick question: Do you consider non-PhD candidates with competition-proven protein design experience?
- 50 proteins designed and ranked #386/681 (Nipah Binder Competition - ProteinBase) with only laptop.
- #207/337 globally (ARC Virtual Cell Challenge)
- Drug discovery platforms: 1,192+ molecules validated
I know I don't have the PhD, but I've built what you're building. Thoughts?
You need builders who ship. I ship. Let's talk.
English

@AllThingsApx proceedings.neurips.cc/paper_files/pa… <- from NeurIPS 2023.
Also check out openreview.net/forum?id=GZ7gw… <- from ICML this year
English

Quietly, this paper reset the trajectory of protein language modeling earlier this month.
Retrieval-at-inference (MSAs) + pair-representation beats 10–100× larger single-sequence LMs on contacts, PPIs, variant effects.
Not just a benchmark win - a blueprint: query-biased MSAs, explicit pair geometry, stress-tested reliability.

English

PoET-2 is open source on github: github.com/OpenProteinAI/…
Thanks to the @openprotein team!
English

Our preprint on sequence-to-property learning and zero-shot fitness prediction with PoET-2 is live: arxiv.org/abs/2508.04724
English
tbepler ری ٹویٹ کیا

Understanding Protein Function with a Multimodal Retrieval-Augmented Foundation Model
1. PoET-2, a new protein language model, achieves state-of-the-art performance in predicting the effects of mutations on protein function, especially for challenging cases like insertions/deletions and higher-order mutations. This model combines sequence, structure, and evolutionary information in a novel way to improve protein understanding and design capabilities.
2. The model incorporates a hierarchical transformer encoder and dual decoders with both causal and masked language modeling objectives. This dual training approach allows PoET-2 to excel in both generative tasks (like sequence generation) and bidirectional representation learning, making it versatile for various protein-related tasks.
3. PoET-2 leverages retrieval augmentation, which enables it to learn from context and incorporate new sequences not present in the original training data. This feature enhances its ability to adapt to different protein families and their specific evolutionary constraints, leading to more accurate predictions.
4. In zero-shot variant effect prediction, PoET-2 outperforms previous models significantly, especially on datasets involving multiple mutations and indels. It also shows superior performance in supervised settings with limited data, demonstrating excellent data efficiency and generalization ability.
5. The model's architecture includes a structure-based attention bias mechanism, which integrates structural information into the attention operations. This enhances the model's ability to capture 3D structural relationships, contributing to its improved performance in tasks related to protein structure and function.
6. PoET-2 is compact, with only 182 million parameters, making it efficient and scalable. Despite its smaller size, it matches or exceeds the performance of much larger models, highlighting its efficiency and practicality for real-world applications in protein engineering and design.
7. The authors demonstrate PoET-2's effectiveness across various benchmarks, including deep mutational scanning and clinical datasets. The model's ability to predict the fitness effects of mutations accurately can accelerate the development of new therapeutics and enhance our understanding of disease mechanisms.
📜Paper: arxiv.org/abs/2508.04724
#ProteinEngineering #AIinBiology #MachineLearning #ProteinFunction #MutationPrediction

English

PoET-2 (and more) is available now on the @openprotein cloud.
We also open sourced PoET-2 on github: github.com/OpenProteinAI/…
Thanks to @timt1630 and the rest of the @openprotein team!
English

Learn more in our pre-print: arxiv.org/abs/2508.04724
or blog post: openprotein.ai/a-multimodal-f…
English

If you're interested in the PLM used here, check out PoET-2 from @openprotein! It allows controllable protein generation with sequence (homologue) and optional structure conditioning and is available right now. 🦾
Boris Power@BorisMPower
At @OpenAI, we believe that AI can accelerate science and drug discovery. An exciting example is our work with @RetroBiosciences, where a custom model designed improved variants of the Nobel-prize winning Yamanaka proteins. Today we published a closer look at the breakthrough. ⬇️
English

Nice article about @lab_berger 's new work on interpreting PLMs!
news.mit.edu/2025/researche…
English
tbepler ری ٹویٹ کیا

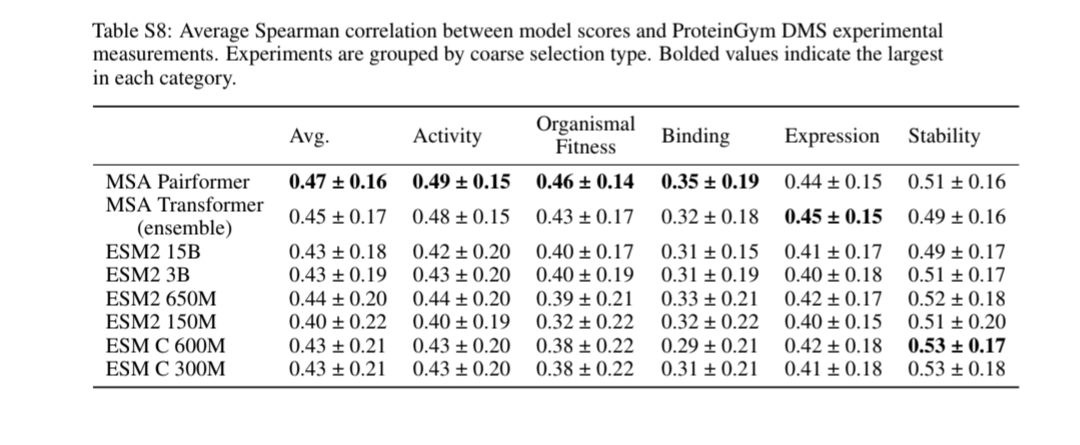
@sokrypton @ferruz_noelia @ruben_weitzman @NotinPascal @yoakiyama I got the 0.45 number from this table
English

@tbepler1 @ferruz_noelia @ruben_weitzman @NotinPascal @yoakiyama See SI. Even when we break it down by category type, MSA pairformer appears to do better than MSA transformer and ESM2.
English

I’ve thoroughly enjoyed reading two (VERY!) recent papers that model protein sequences by retrieving evolutionary information (dynamically) at inference time, and there's a lot to unpack!
[1] arxiv.org/abs/2506.08954
[2] biorxiv.org/content/10.110…
(1/n)
English

@sokrypton @ferruz_noelia @ruben_weitzman @NotinPascal @yoakiyama I'm not critiquing your internal comparison here, but rather that your Avg. is not comparable with the main number on the proteingym benchmark since it isn't aggregated the same way
English

@sokrypton @ferruz_noelia @ruben_weitzman @NotinPascal @yoakiyama The average spearman in proteingym is a macro average over the spearman correlations by function since some functions have significantly more datasets
English

@tbepler1 @ferruz_noelia @ruben_weitzman @NotinPascal @yoakiyama Can you expand on this? What do you mean by properly averaging?
I believe we computed the values the same across the methods we show, so it should be a fair comparison. Unless we made a mistake?
English
