Finn
243 posts


Finn
@finnlueth
building @protzilla at @fdotinc. (ᴅʀᴜɢs, ᴘʀᴏᴛᴇɪɴs) × ᴍʟ. bioinformatics @TU_Muenchen. researcher @rostlab. prev @iGEM_Munich & steineggerlab. 🇺🇸🇩🇪.

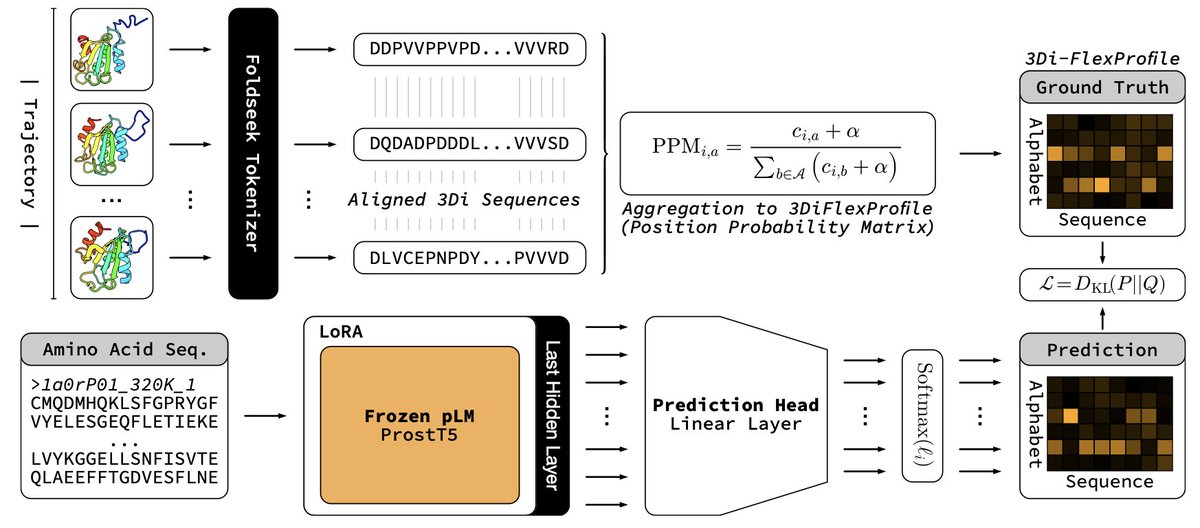

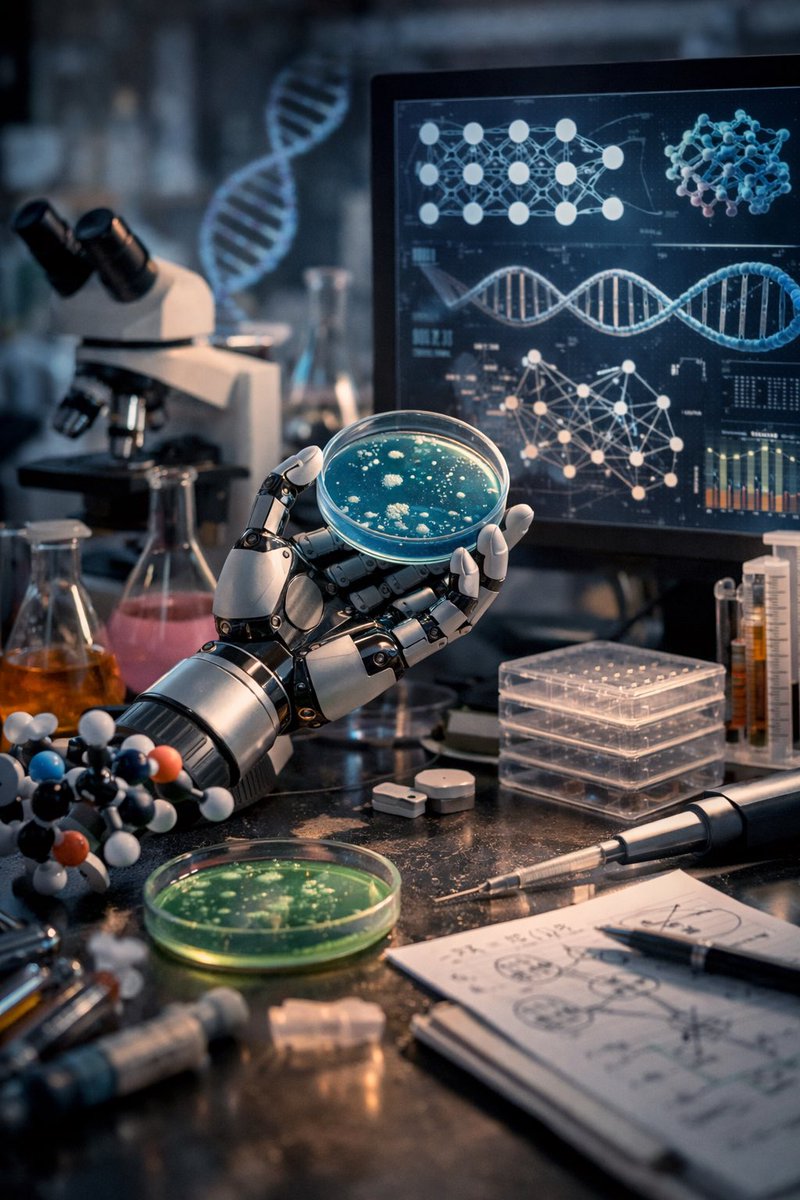
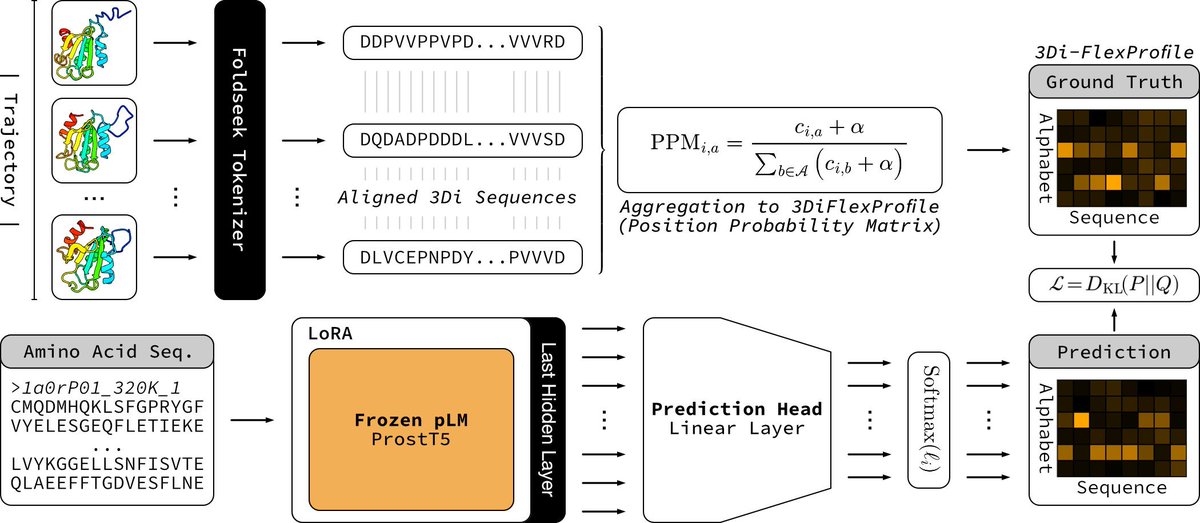
My bachelor's thesis (Protein Language Modeling beyond static folds reveals sequence-encoded flexibility) is now a preprint. ProtProfileMD is a fine-tune of ProstT5 that learned per-residue 3Di probability profiles generated from mdCATH molecular dynamics trajectories. The probability profiles recovered flexibility signals and boosted remote homology detection. Thanks to my supervisors, advisors, and collaborators @HeinzingerM @BurkhardRost, Steinegger Lab, and @rostlab for making it possible.






The results of the Nipah Protein Design Competition are out! 🧬 1200 proteins experimentally validated (3x more than last year) 📈 99 novel binders against the target protein (a challenging tetramer with little prior work) 💪 26 single digit nM or better binders, with the best ones at single-digit picomolar affinity! All data now available open-source on Proteinbase! Let's take a look at the results ⬇️


In Packy McCormick's recent deep dive on a16z, he writes, “What a16z aims to do is provide legitimacy and power [for startups]”. a16z cofounders Marc Andreessen and Ben Horowitz have been building the venture firm to provide entrepreneurs with legitimacy and power for almost two decades. In this conversation, they join Packy and a16z GP Erik Torenberg to cover how they did it and the worldview behind a16z, including: - How a16z compounds reputation - How the media ecosystem has changed since a16z began & how a16z has adapted - How a16z is structured to put entrepreneurs first and enable them to win - a16z's culture document and how written culture shapes people's actions - How to size markets that will grow exponentially because of technology - Why there are so many great Zoomer founders and much more. 0:00 Introduction 00:46 How the media ecosystem is changing 4:20 Substack 6:28 Supply-driven markets and new content creation 10:09 Databricks 13:58 Demand for great content 18:49 Market sizing 22:37 Turning inventors into confident CEOs 27:29 Building dreams 30:46 Compounding reputation 40:39 a16z team structure 46:01 Why intangibles matter more than ever 48:17 Original thinkers with charisma 50:06 Zoomers @pmarca @bhorowitz @packyM @eriktorenberg Not an offer or solicitation. None of the information herein should be taken as investment advice; Some of the companies mentioned are portfolio companies of a16z. Please see a16z.com/disclosures/ for more information. A list of investments made by a16z is available at a16z.com/portfolio.













