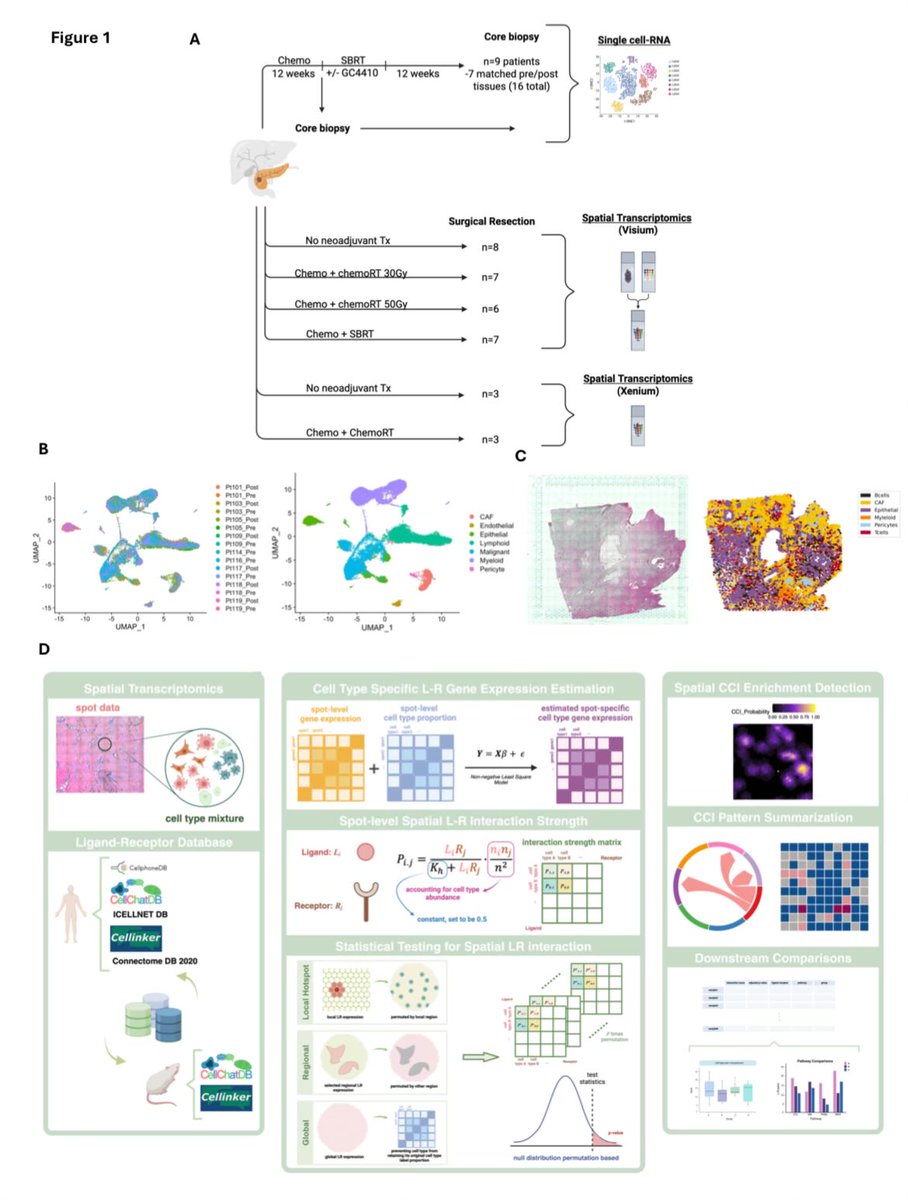
@LyssiotisLab Agree! This is nuts! Combining synthetic biology with in vivo understanding to give an intervention like this!
English
Bednar lab
351 posts


@BednarLab
Pancreatic cancer research, epigenetics, tumor immunology, tumor evolution, surgeons doing basic science, UM PanTErA, #NewPISlack, personal Twitter: @Fil_Bednar



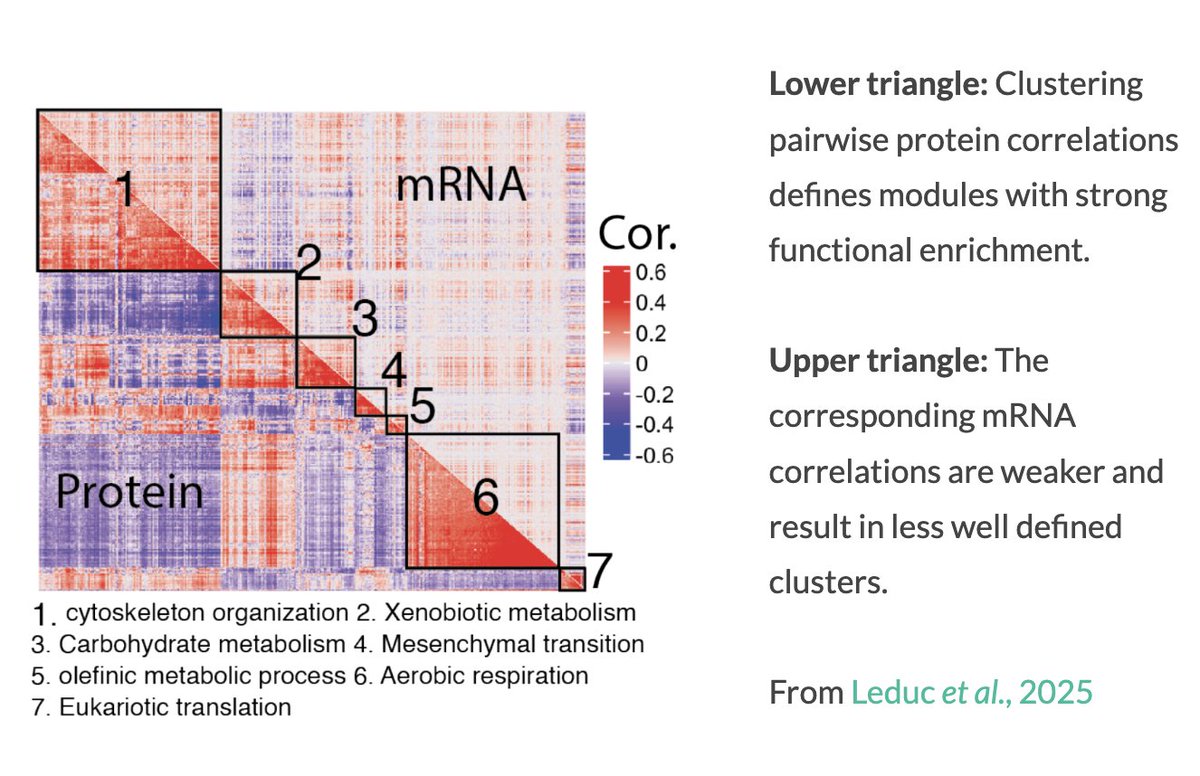



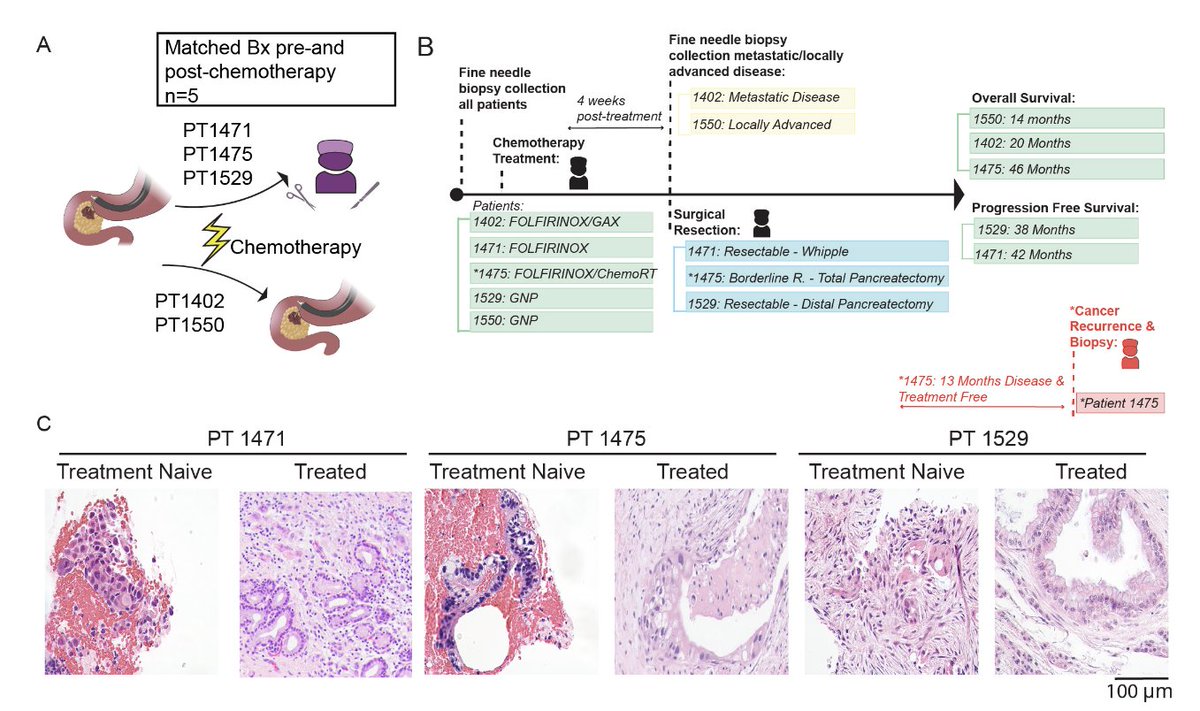



The first rule of Data Science - if your p-value is less than 1 over the number of atoms in the universe, you're using the wrong model.























Submit an abstract by July 15 for the AACR Conference on Advances in Pancreatic Cancer Research (Sept 28-Oct 1; Boston), chaired by Andrew J. Aguirre, @EdnaCukierman, Susan Tsai, David Tuveson, and Laura DeLong Wood. brnw.ch/21wTK6Y #AACRpan25 @isteaus @lauradelongwood