Anush Chiappino-Pepe
2.6K posts


Anush Chiappino-Pepe
@AnushChP
Engineering codes of life 4 cellular insights + chem/fx/tech w #Syn/CompBio | PostDoc GMChurch @Harvard/Wyss, GStephanopoulos @MIT | #BioChemE #GCE #Metabolism






30 Underrated Ideas in Biotech (#6) Engineered organisms are everywhere, but they are usually "contained." Engineered cells make many of our medicines, for example, but they are isolated to bioreactors. Some of the more "sci-fi" visions of biotechnology -- like cleaning up an oil spill that spans many square miles of ocean, or using microbes at scale to degrade plastics -- require that we first release engineered organisms *without* containment. Of course, there are strict regulations preventing this. Getting approval to release a microbe into the wild, without containment, is extremely difficult. The regulatory pathway for genetically-engineered microbes is divided across the EPA, USDA, and FDA. Each agency has jurisdiction depending on the intended use. Biopesticides fall to the EPA, while other agricultural products go to the USDA, and ingestible microbes fall under the province of the FDA. Anything that DOES NOT easily fit into these categories, such as environmental biosensors or microbes engineered to clean up oil spills in the ocean, is instead lumped under the EPA’s Toxic Substances Control Act, or TSCA. Regardless of your stance on engineered microbes, TSCA is not a good regulation. It regulates genetically engineered microbes based on the METHOD by which they were engineered, rather than the product itself. Any microbe containing DNA from another genus — say, moving a gene from Escherichia coli into Pseudomonas putida — is flagged by TSCA and unlikely to get approval, even if researchers can prove that the product is safe. So whereas engineered crops can get approval if researchers provide compelling data from field trials and safety tests, there is no such option for the organisms that get lumped into TSCA. More than 200 TSCA submissions were filed between 1987 and 2018; none of those submissions have led to a commercialized product. Therefore, most companies that want to release a genetically-engineered microbe find ways to skirt around it. Pivot Bio, for example, sells genetically-engineered microbes that colonize plant roots and convert atmospheric nitrogen (N₂) into ammonia (NH₃), a chemical form that plants can use. This reduces the amount of fertilizer needed for a field (a good thing!) thus decreasing the leaching of fertilizer byproducts into water. Pivot Bio sidestepped some regulatory hurdles by avoiding the transfer of genes from one species to another; they simply remodeled their organism’s existing genome. The company still has to get USDA approval to ship its product across state lines, but that is a simpler regulatory hurdle to get over. If we want to see more "sci-fi" biotech in the world, we should rewrite TSCA and build a clear path for companies to file for approval based on actual field trials & safety testing.

I’m excited to announce that the application for the 2026 edition of our @medialab and global synthetic biology course “How to Grow (Almost) Anything” is now open! Application link in thread 👇



Struggling with weight loss? GLP-1 treatments backed by science make it easy. Affordable, effective, and delivered to your door. Start your journey today!




New episode w @AdamMarblestone on what the brain's secret sauce is: how do we learn so much from so little? Also, the answer to Ilya’s question: how does the genome encode desires for high level concepts that are only seen during lifetime? Turns out, they’re deeply connected questions. Timestamps 0:00:00 – The brain’s secret sauce is the reward functions, not the architecture 0:22:20 – What the genome actually encodes 0:42:42 – What kind of RL is the brain doing? 0:50:31 – Is biological hardware a limitation or an advantage? 1:03:59 – Why we need to map the human brain 1:23:28 – What value will automating math have? 1:38:18 – Architecture of the brain Look up Dwarkesh Podcast on YouTube, Apple Podcasts, Spotify, etc. Enjoy!

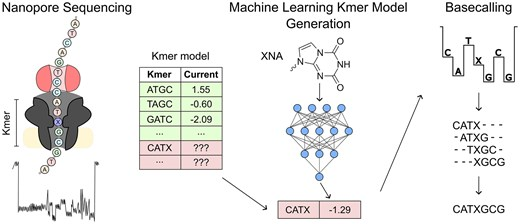

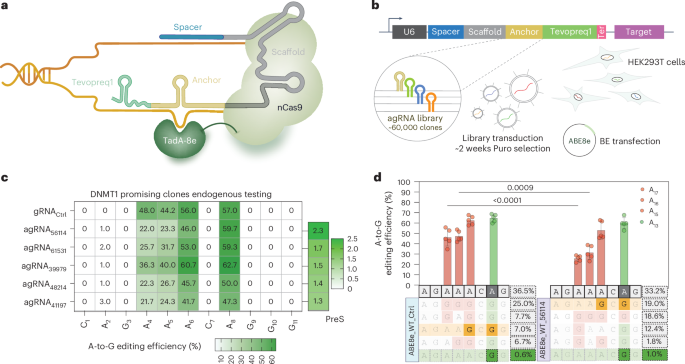

Our new paper is now online in @NatureBiotech. We use gRNA engineering, phage-assisted non-continuous evolution (PANCE), and protein language models to improve the precision of adenine base editing. @harvardmed @geochurch nature.com/articles/s4158…


