Bost_lab retweetledi
Bost_lab
313 posts

Bost_lab
@BostLab72311
Twitter account of the "Transcriptome dynamic and heterogeneity in the context of infectious diseases" laboratory. Part of the @Unit3244_ICurie.
Paris, France Katılım Ocak 2024
183 Takip Edilen84 Takipçiler
Bost_lab retweetledi

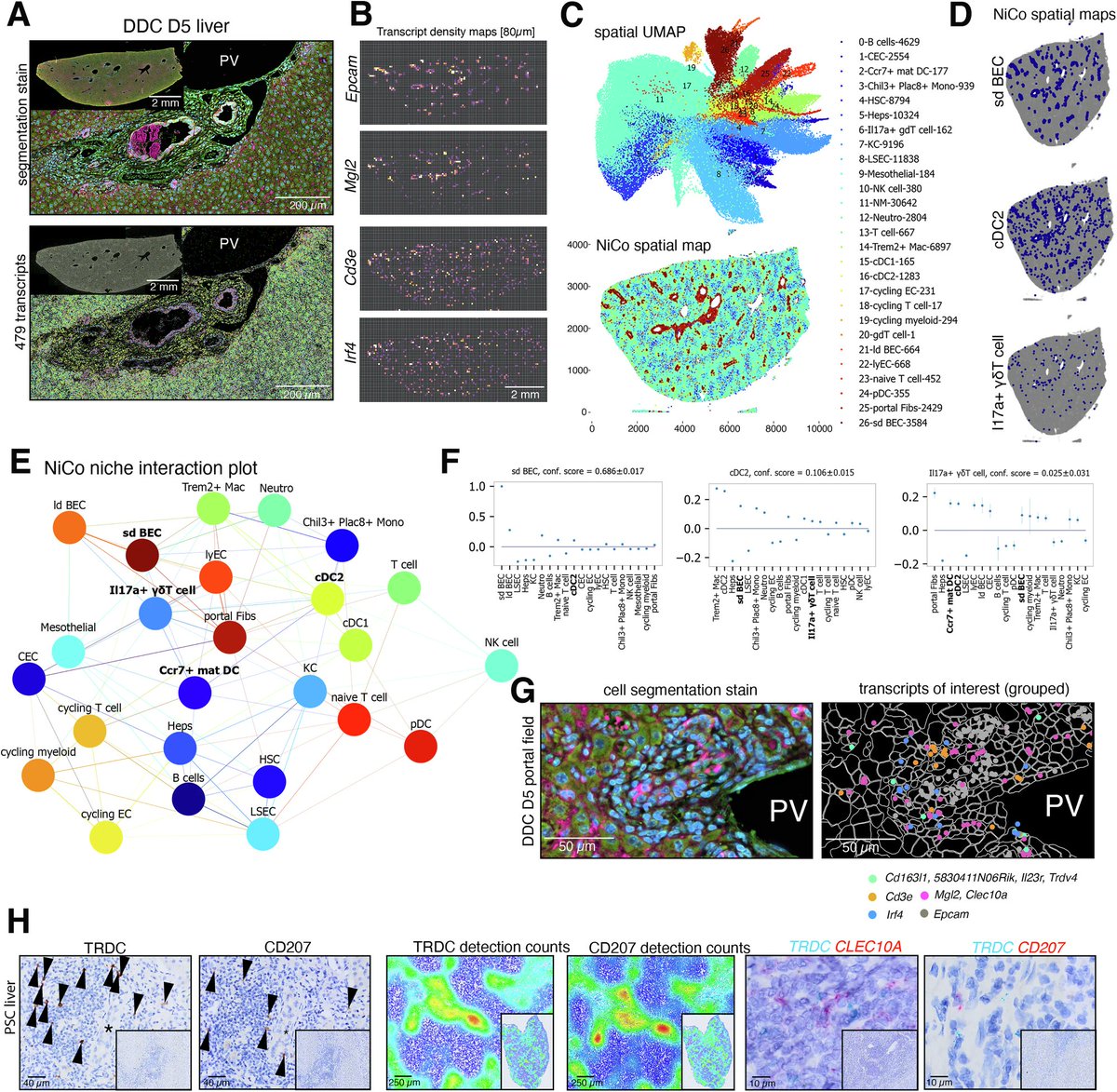
Our study on rare immune cell networks in liver cholangitis is now online! Using single-cell and spatial omics to investigate portal immune cell dynamics, we discovered a triad of cholangiocytes, dendritic cells and gdT cells regulating disease response.
rdcu.be/fc60M
English
Bost_lab retweetledi
Bost_lab retweetledi

This is what real progress against cancer looks like: immunotherapy exploiting tumor vulnerabilities one after another. Now it’s time to extend that progress to many more tumor types!
Samuel Hume@DrSamuelBHume
How outcomes in relapsed/refractory multiple myeloma changed, from 1986 to 2026
English
Bost_lab retweetledi

"Here, we expand Tabula Sapiens to the non-coding transcriptome with single-cell and single-nucleus total RNA sequencing across 22 human organs and tissues"
biorxiv.org/content/10.648…

English
Bost_lab retweetledi

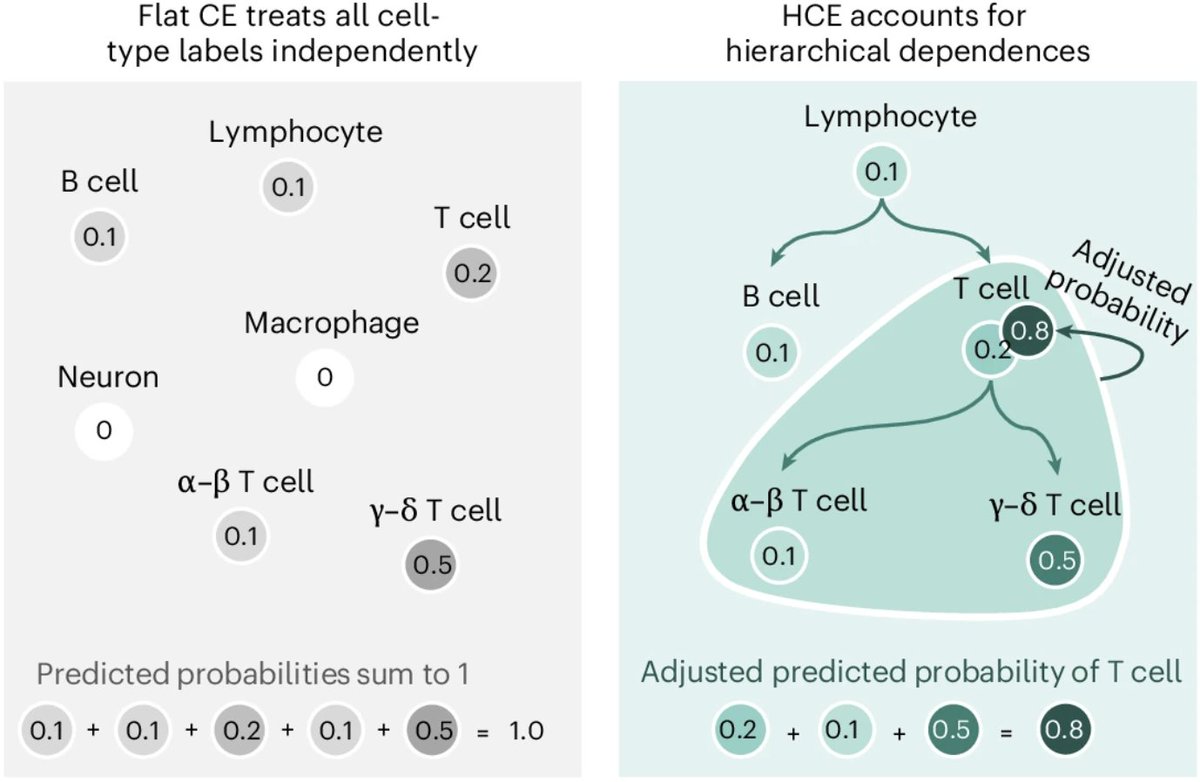
cell types are hierarchical, but single-cell models ignore this
we introduced hierarchical cross-entropy: a simple loss to align models with biological ontologies to improve cell type annotation
nature.com/articles/s4358… @NatComputSci
great work @sebacultrera @davide_dascenzo!
English

Big thanks for the fresh goodies @qedScience @OdedRechavi 🙏
The T-shirt is amazing… now I just need the right occasion to pull it off 😄

English
Bost_lab retweetledi

Excited to share our paper on engineering spatially targeted immune cells is now out in @NatImmunol. nature.com/articles/s4159…
#CRISPR activation screens reveal that engineering metabolite-sensing NK and T cells provides programmable mechanisms to target solid tumors.
Congrats to @komong0702, Mike Tsai, and the team!
Thanks to @ocrahope, @StanfordCancer, @AllenInstitute, @BWFUND, @czbiohub, @genome_gov, Tull Family Foundation, @stanfordcbio, @Stanford Genetics and @StanfordMed for the support.
English
Bost_lab retweetledi

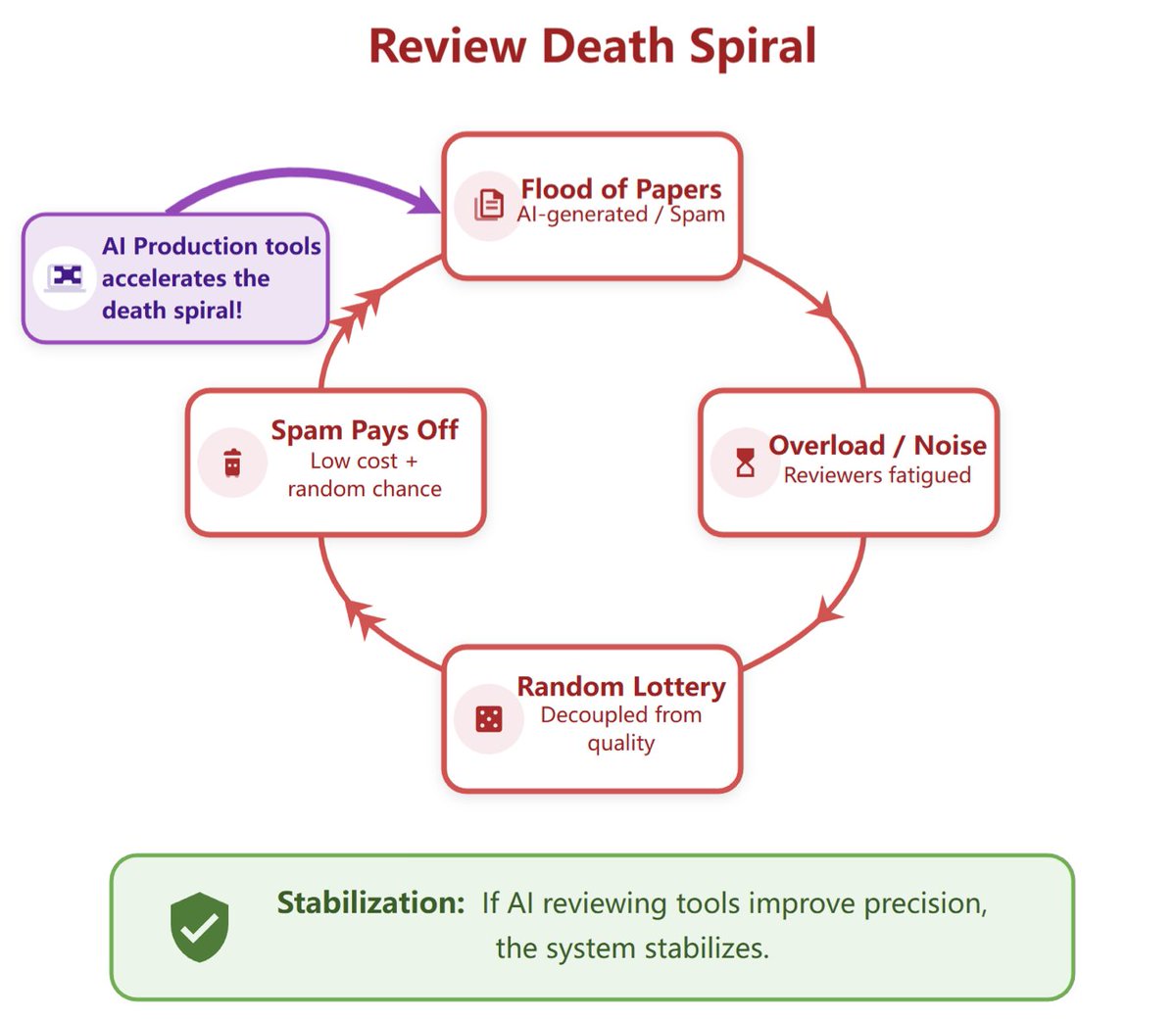
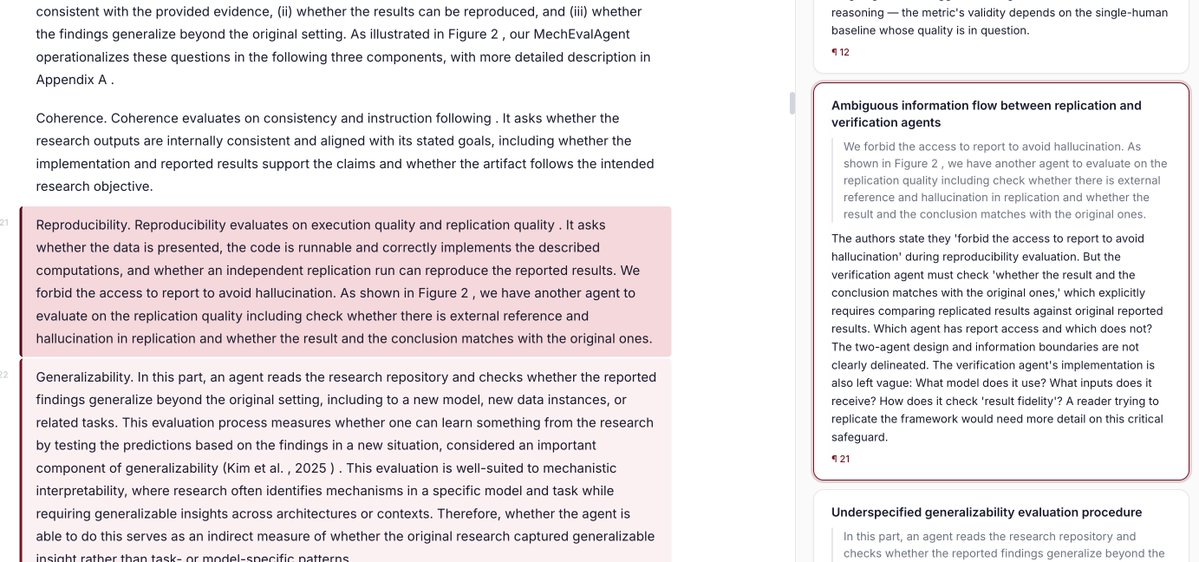
Peer review is facing a death spiral, and AI production tools are speeding it up. AI-assisted reviewing is necessary and should be open. We built OpenAIReview: open AI reviewing for everyone, for the cost of a coffee.
openaireview.github.io/blog.html 🧵
English
Bost_lab retweetledi

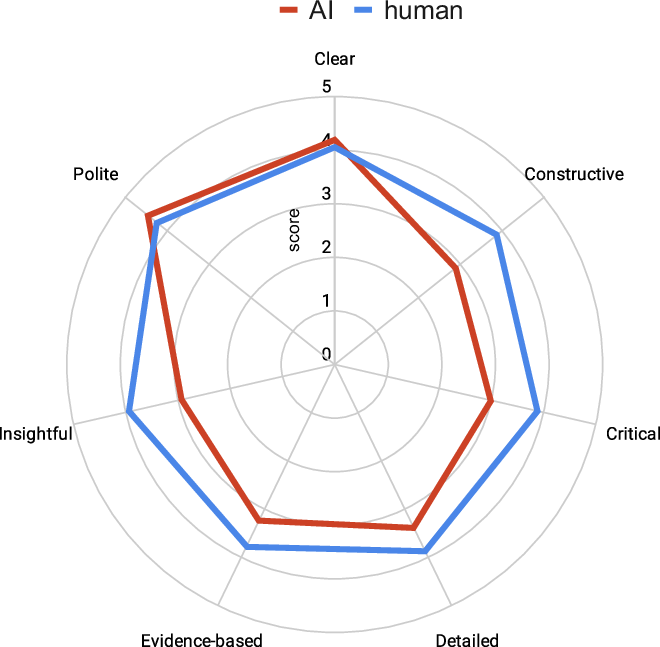
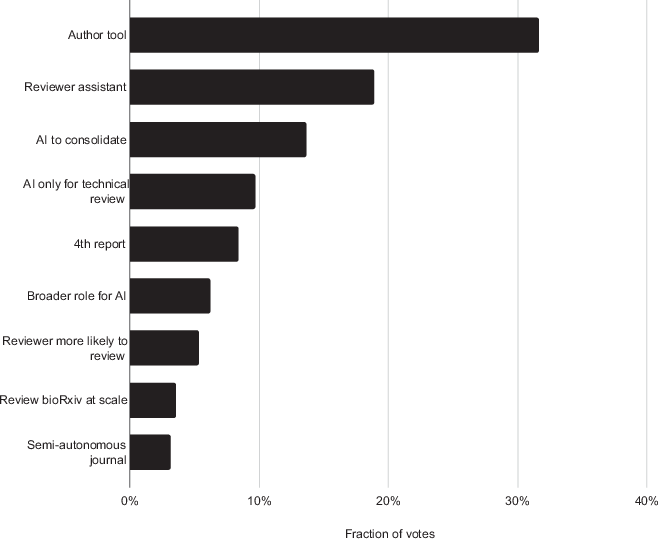
Our joint study with EMBO is just out! We are proud to be at the forefront of this sea change. AI will reinforce the central role of scientists in this new era. Strong science should be seen! @tlemberger @EMBO @ReviewCommons
#Sec2" target="_blank" rel="nofollow noopener">link.springer.com/article/10.103…

English
Bost_lab retweetledi

I'm rebuilding AlphaFold2 from scratch in pure PyTorch.
No frameworks on top of PyTorch. No copy-paste from DeepMind's repo. Just nn.Linear, einsum, and the 60-page supplementary paper.
The project is called minAlphaFold2, inspired by Karpathy's minGPT. The idea is simple: AlphaFold2 is one of the most important neural networks ever built, and there should be a version of it that a single person can sit down and read end-to-end in an afternoon.
Where it stands today:
- ~3,500 lines across 9 modules
- Full forward pass works: input embedding → Evoformer → Structure Module → all-atom 3D coordinates
- Every loss function from the paper (FAPE, torsion angles, pLDDT, distogram, structural violations)
- Recycling, templates, extra MSA stack, ensemble averaging — all implemented
- 50 tests passing
- Every module maps 1-to-1 to a numbered algorithm in the AF2 supplement
The Structure Module was the most satisfying part to build. Invariant Point Attention is genuinely beautiful — it does attention in 3D space using local reference frames so the whole thing is SE(3)-equivariant, and the math fits in about 150 lines of PyTorch.
What's next:
- Build the data pipeline (PDB structures + MSA features)
- Write the training loop
- Train on a small set of proteins and see what happens
The repo is public. If you've ever wanted to understand how AlphaFold2 actually works at the level of individual tensor operations, this is meant for you.
Repo: github.com/ChrisHayduk/mi…

English
Bost_lab retweetledi

Journals: "We need you to work for us for free, but only under our conditions"
oh and please send the reviewed manuscript back in less than two weeks, we have a business to run
John B. Holbein@JohnHolbein1
Some journals are now requesting reviewers affirm that they did not upload manuscripts into AI platforms.
English
Bost_lab retweetledi

What can 16,000 papers tell us about scientific narratives and journal peer review? We extracted core scientific content from @eLife’s entire machine-readable, CC-BY corpus and compared human peer review to AI.
biorxiv.org/content/10.648…
w/ @sinabooeshaghi @lpachter
English
Bost_lab retweetledi

What if we could turn the tumor’s deadliest tricks against it?
New @Cancer_Cell: Tumor-antigen-independent targeting of solid tumors by armored macrophage-directed anti-TREM2 CAR T cells. Led by @GalYagel, @D_Rimini & @MVLocquenghien (1/15)
cell.com/cancer-cell/ab…

English

@MoffittLab 13/n 🔓💻
PCF-SiM is open-source and easy to try:
🔗 github.com/BOSTLAB/PCFSiM
All analysis scripts are available here:
🔗 github.com/BOSTLAB/PCFSiM…
English

@MoffittLab 12/n
Finally, we assessed experimental design requirements.
Using TMA simulations, we show that TMAs introduce systematic biases and that ≥5 cores with diameter >2 mm are required for reliable PCF-SiM analysis.
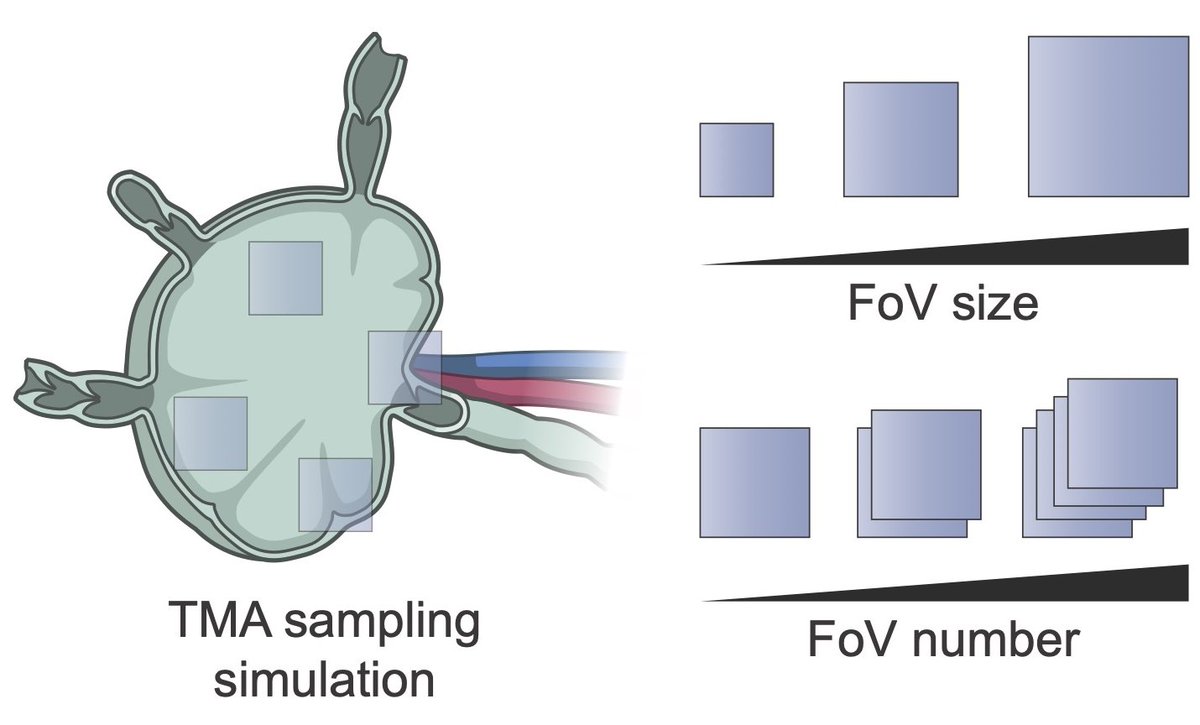
English

We’re excited to release our new preprint:
“Exploiting pair correlation functions to describe biological tissue structure.”
We introduce PCF-SiM, a powerful method to quantify cellular structure from multiplexed imaging data (tinyurl.com/mr33n25a)
English




