Celia Lerma Martin
19 posts

Celia Lerma Martin
@Cels121
PhD student at @schirmerlab. Dry and wet lab, interested in spatial transcriptomics. BSc @UABBarcelona & MSc @UPFBarcelona



Our work with @schirmerlab in multiple sclerosis (MS) is out today in @NatureNeuro. We investigated lesion progression and cell-cell communication events in MS using snRNA-seq and spatial transcriptomics 🧠 See below ⬇️ nature.com/articles/s4159…

Our work with @schirmerlab in multiple sclerosis (MS) is out today in @NatureNeuro. We investigated lesion progression and cell-cell communication events in MS using snRNA-seq and spatial transcriptomics 🧠 See below ⬇️ nature.com/articles/s4159…






Our Featured article: Gene regulatory network inference in the era of single-cell multi-omics go.nature.com/3r8TugN #Review by @PauBadiaM, @LornaWessels, @sophia_mullerd, @TrimbourR, @roramirezf94, @RArgelaguet & @saezlab @UniHeidelberg
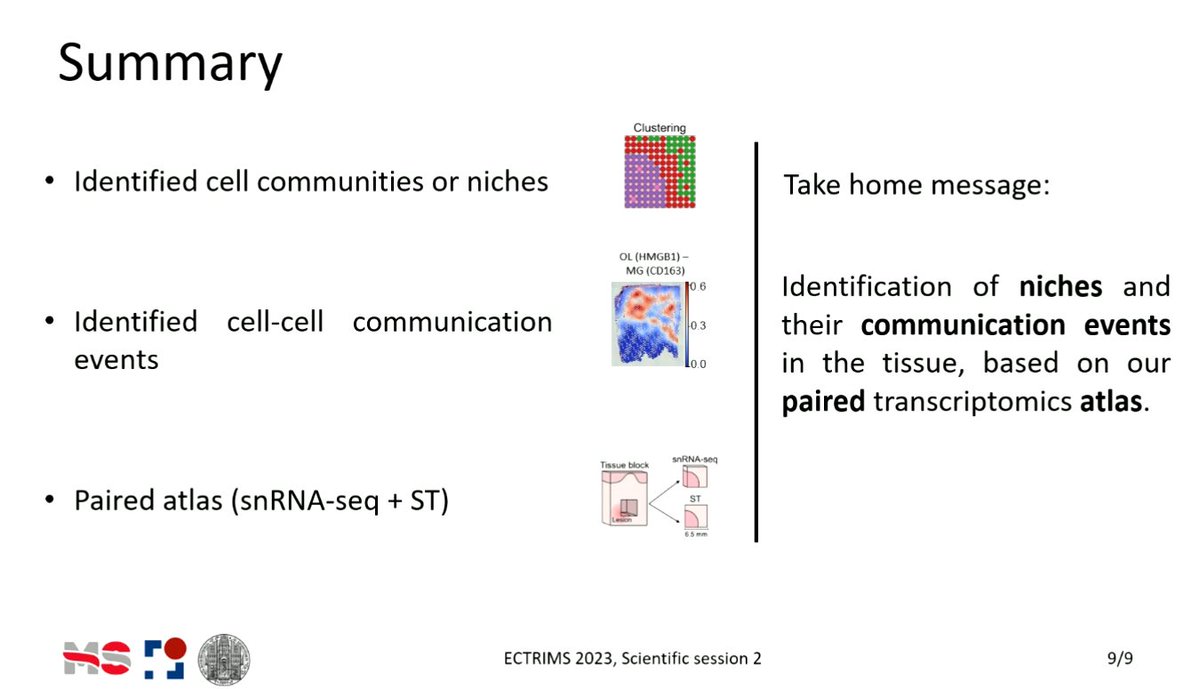





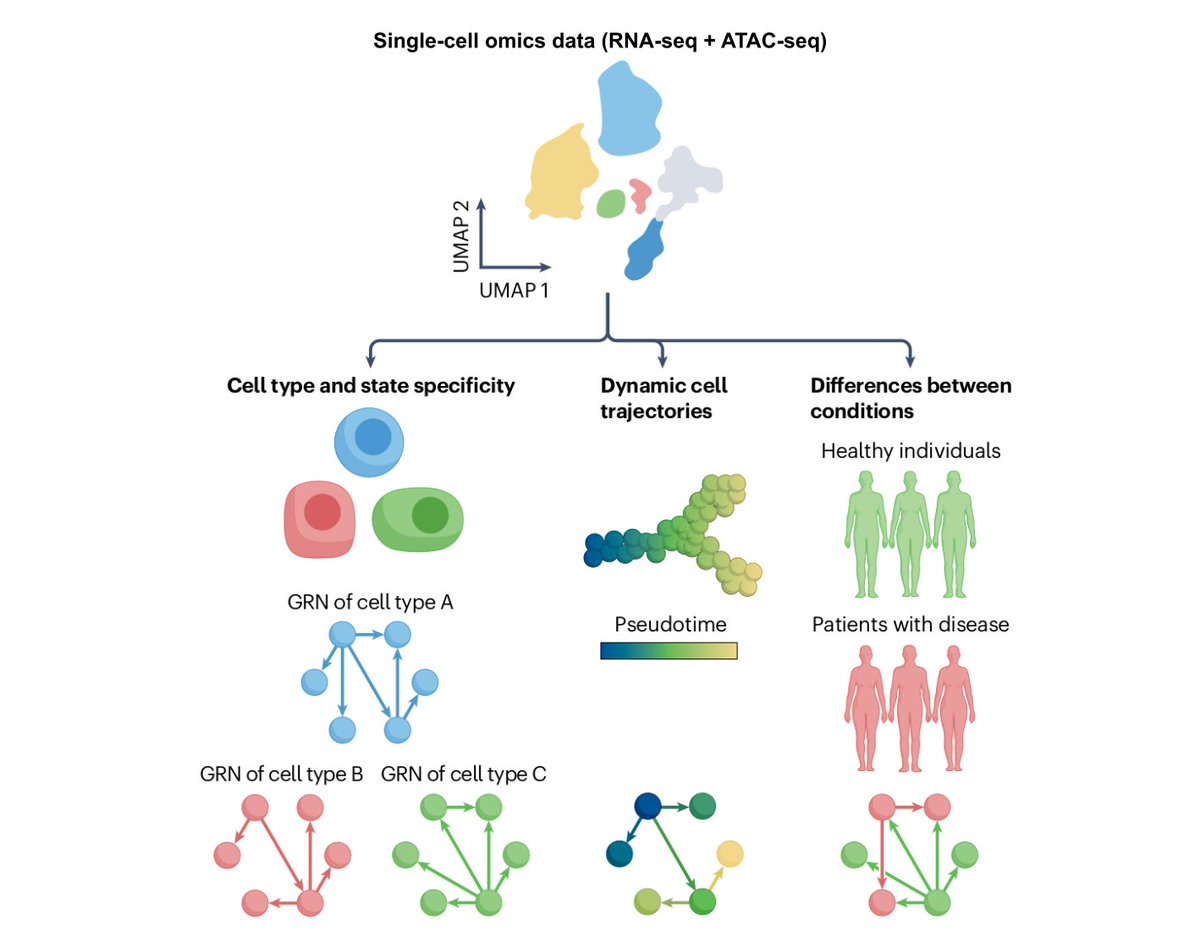


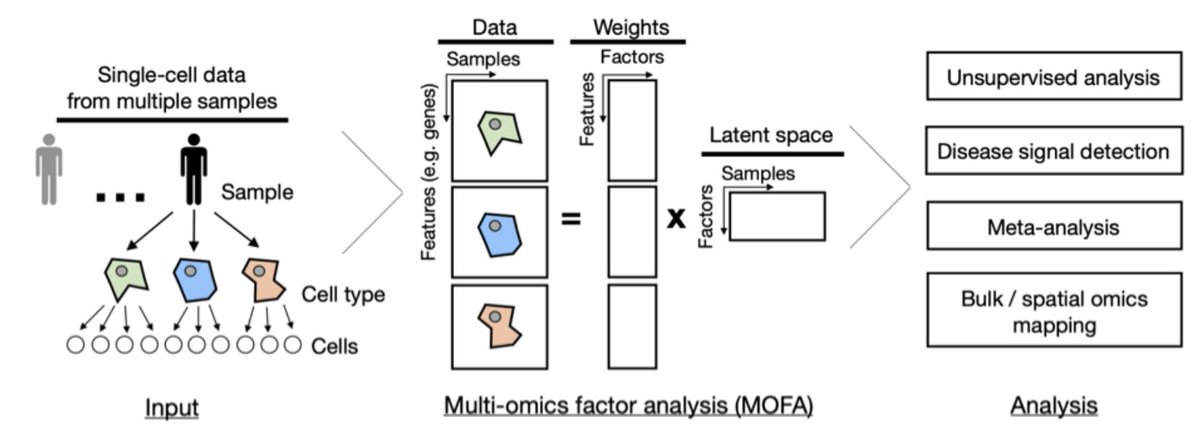



New preprint is out! 🚨 We analyzed the spatial map of multiple sclerosis (MS) lesions using snRNA-seq and spatial transcriptomics 🧠. This was a close collab with @schirmerlab, led by @Cels121 and our @PauBadiaM. See below 👇


Our very much extended and revised multiomics single cell (RNA/ATAC seq) and spatial atlas of human myocardial infarction 🫀✨🚀 is out now @Nature! 🥳 Collaborative effort w/ @rkramann, @vanvanka123, & H Milting’s labs nature.com/articles/s4158…

Great work! We are happy we could help analyze the data with our resources, PROGENy tinyurl.com/2p8razkk and DoRothEA tinyurl.com/3kwn9b7n, and in particular to extract a transcription factor signature present in COVID-19 & other hepatic infections.

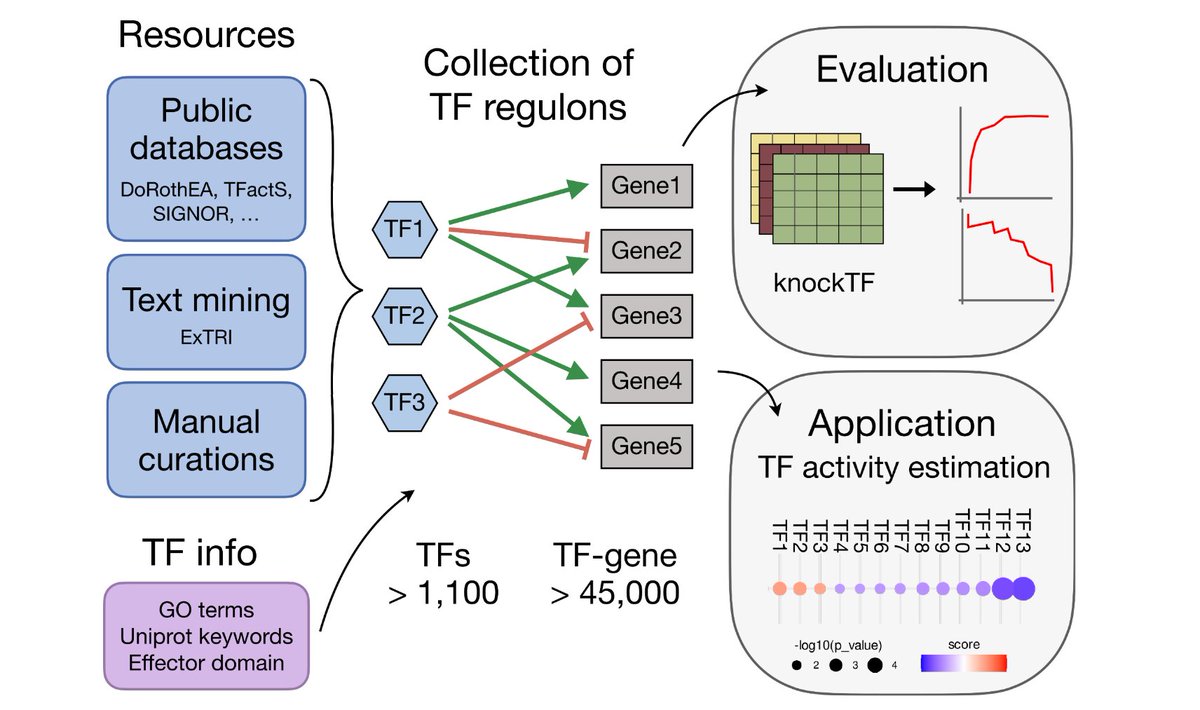
Our updated manuscript describing methods for bioactivity inference (pathways, TFs, etc.) is now published in @BioinfoAdv doi.org/10.1093/bioadv… 🥳 We have been working on a new #Python 🐍 implementation and its integration with the meta-resource @omnipathdb 👇