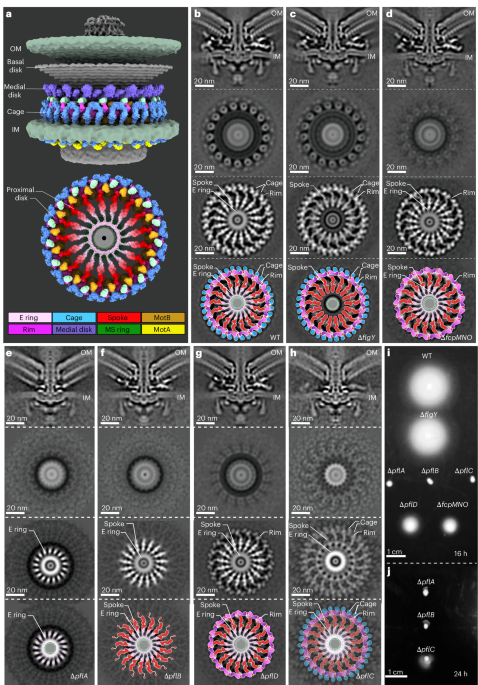
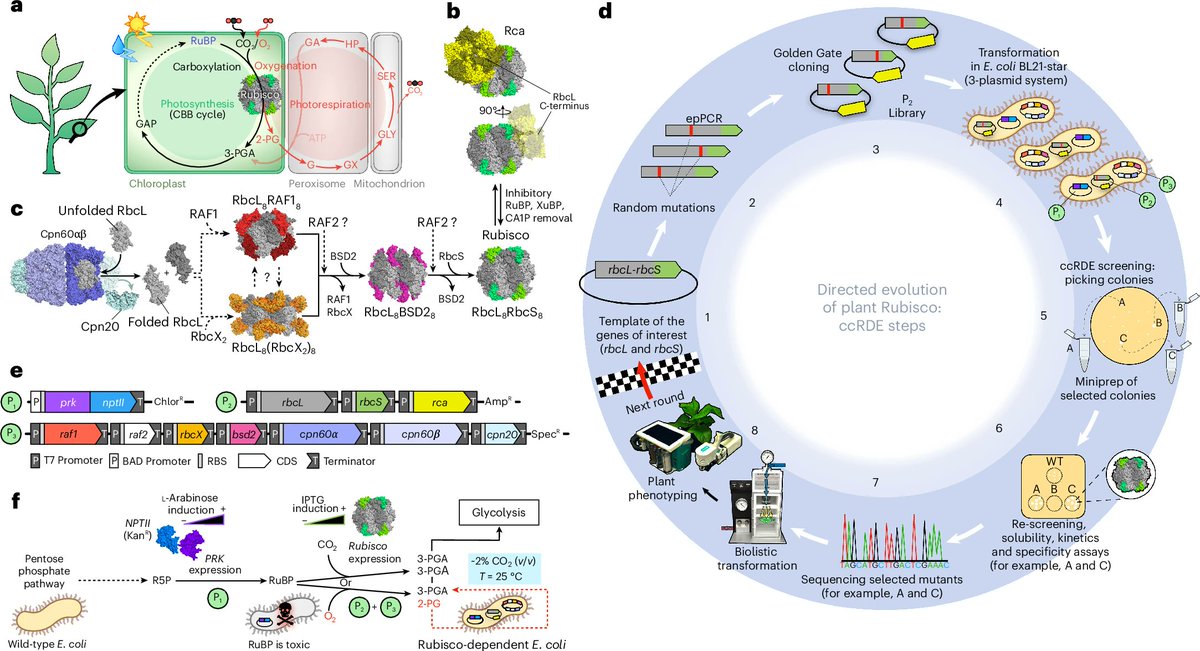
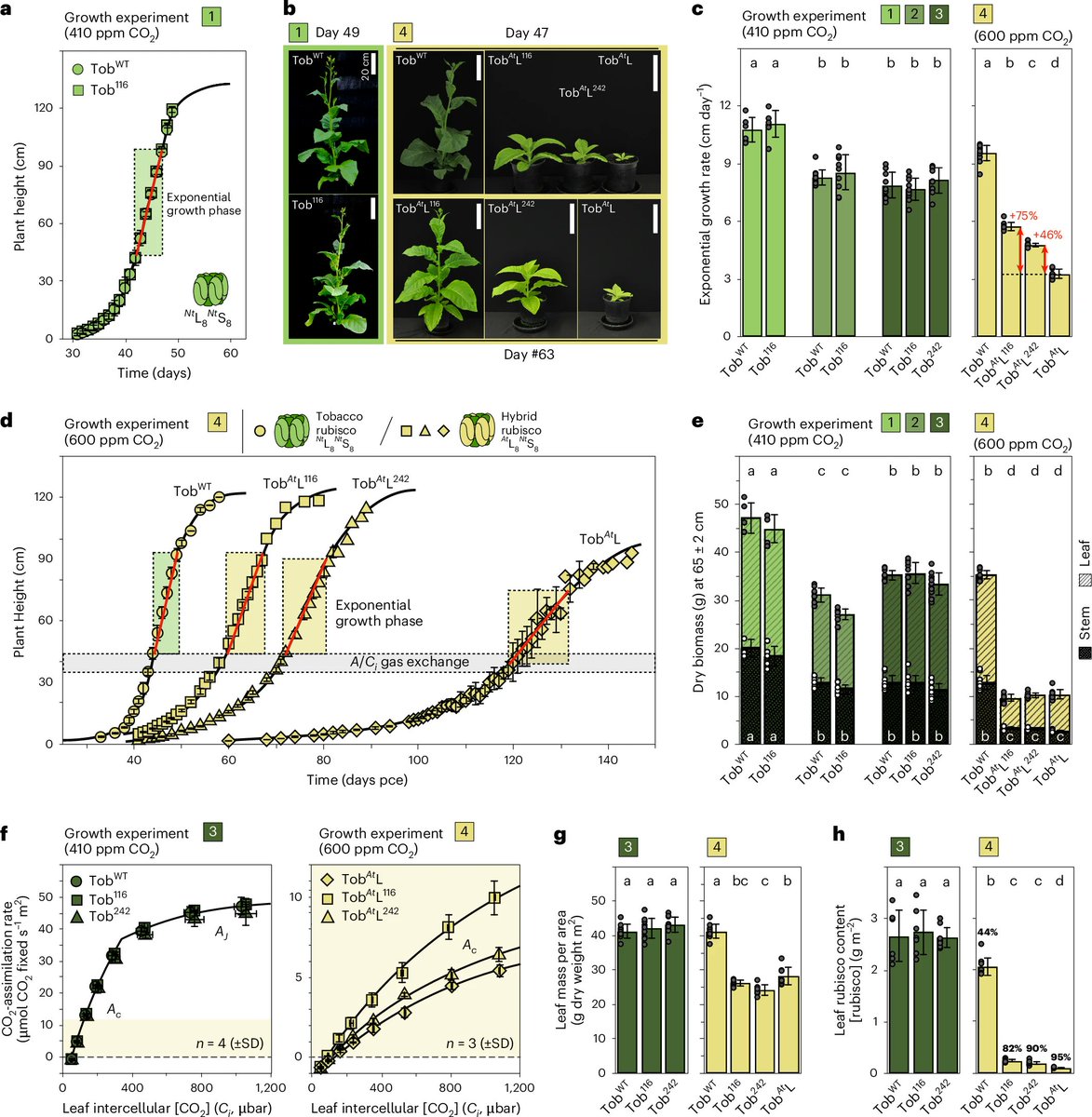
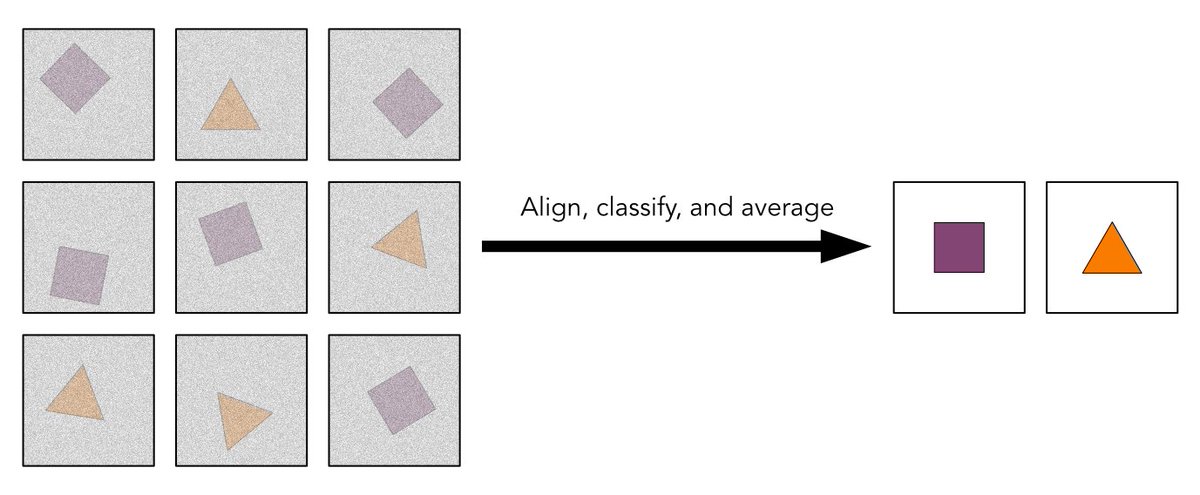
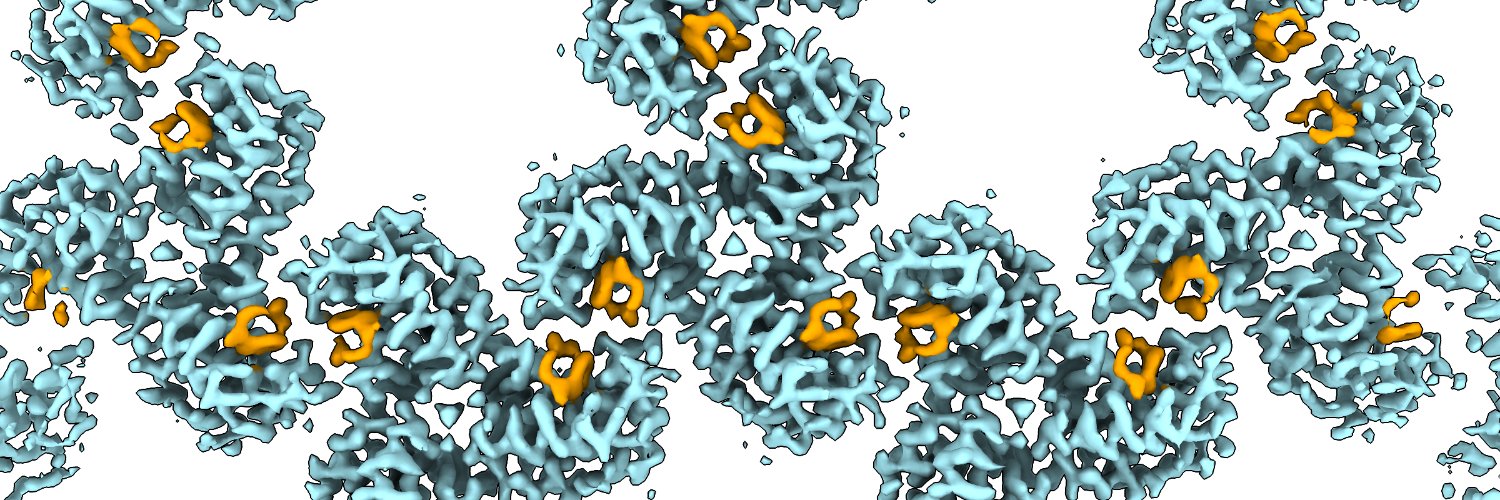
Mature #HIV-1 contains >2000 copies of “spacer peptide 2”, but we didn’t know why. We now see that SP2 binds the matrix protein and triggers the change into its mature arrangement. Summary video from @MargotRiggi :
Dominik Hrebik
1.6K posts

@F4ustus
Currently: Postdoc @BriggsGroup @MPI_Biochem | PhD from @PlevkaLab | #StructuralBiology of #Viruses #HIV | #CryoEM | #Memes Bsky: https://t.co/VOsQ20NTnx


Mature #HIV-1 contains >2000 copies of “spacer peptide 2”, but we didn’t know why. We now see that SP2 binds the matrix protein and triggers the change into its mature arrangement. Summary video from @MargotRiggi :

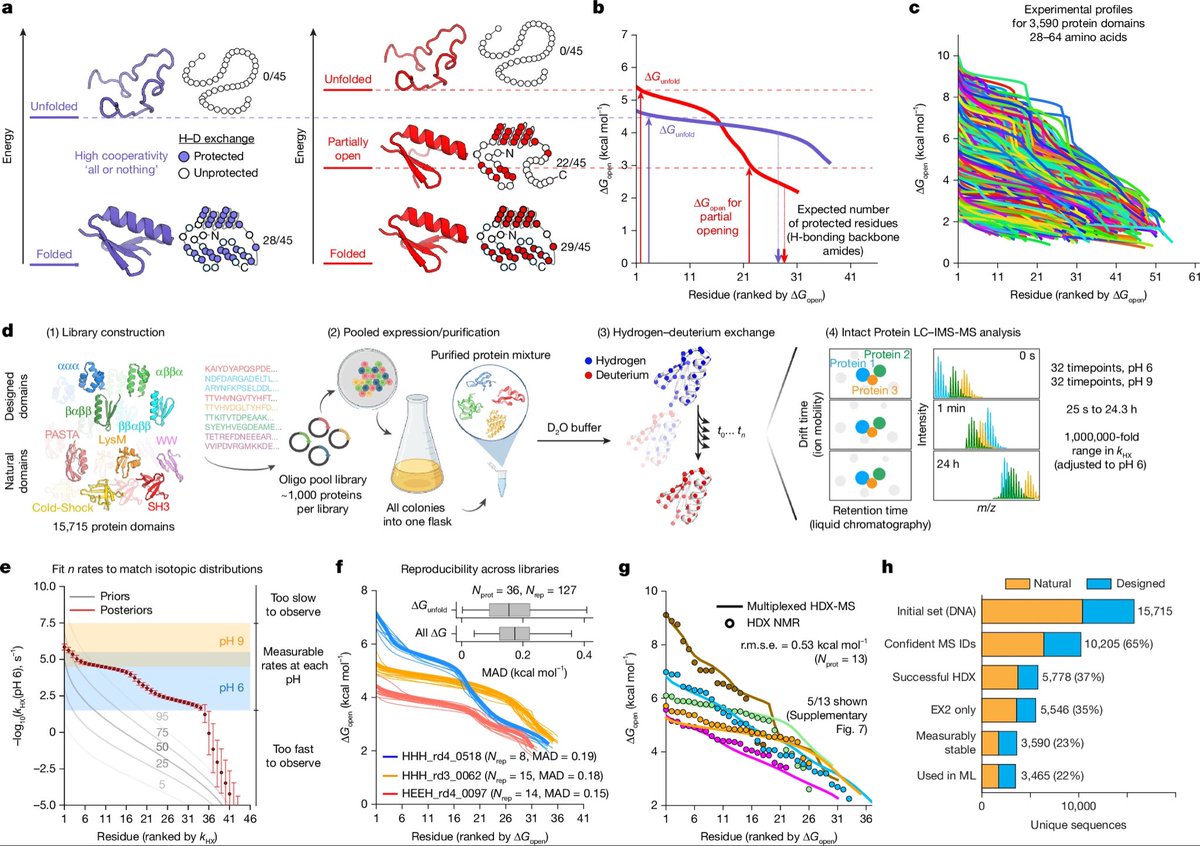
Thrilled to share our new paper where we introduce a multiplexed hydrogen–deuterium exchange MS (mHDX‑MS) method that can measure hundreds of protein domains’ conformational energy landscapes—all in a single experiment! biorxiv.org/content/10.110…




A few years ago, designing an antibody on the computer was extremely difficult. Today, there are several open-source tools which allow anyone to design antibodies from home. Out today: A step-by-step guide to antibody design. By @btnaughton.