Fanying Tang retweetledi

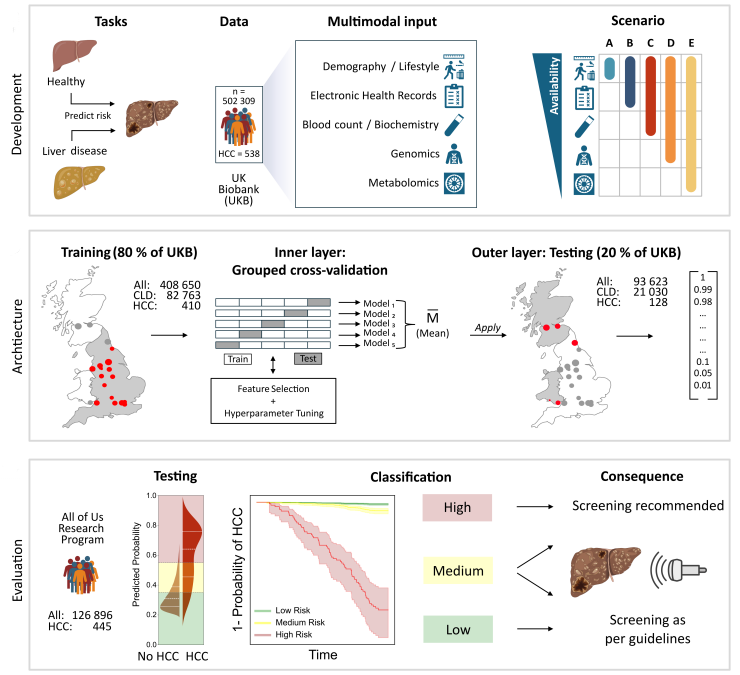
Now online in @CD_AACR: Machine Learning Predicts Hepatocellular Carcinoma Risk from Routine Clinical Data: A Large Population-Based Multicentric Study - by @JClusmann, @jnkath, @C_V_Schneider, and colleagues @RWTH @tudresden_de
Cancer Discovery@CD_AACR
Now online: Machine learning predicts hepatocellular carcinoma risk from routine clinical data: a large population-based multicentric study doi.org/10.1158/2159-8…
English