Gian Marco Visani retweetledi

check out our new manuscript (led by @GMarcoVisani)
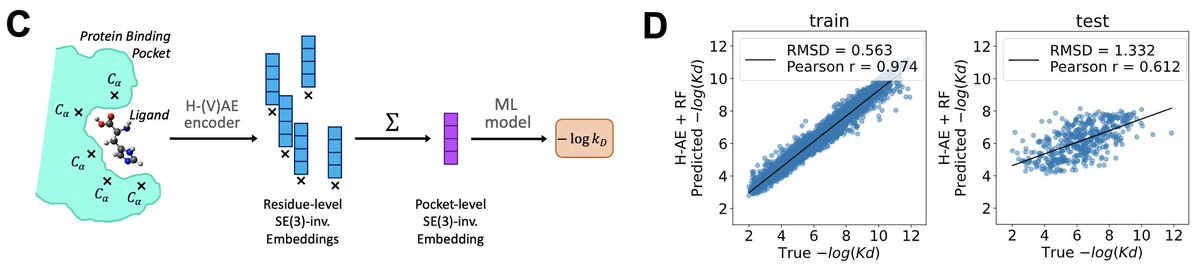
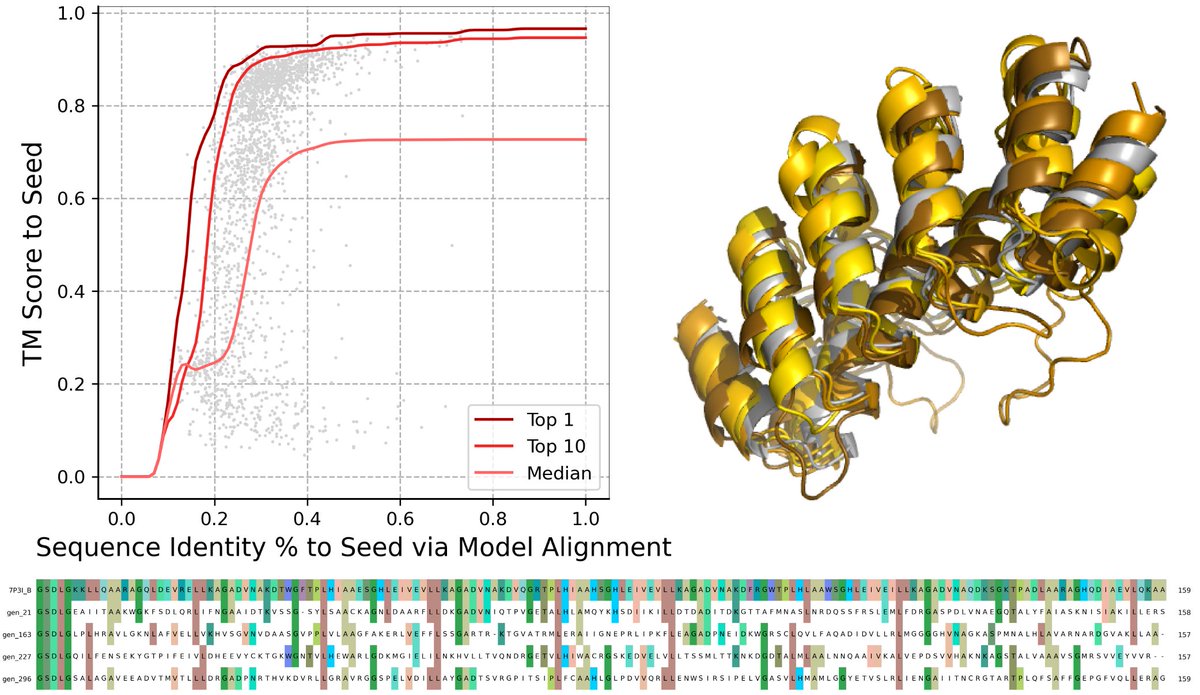
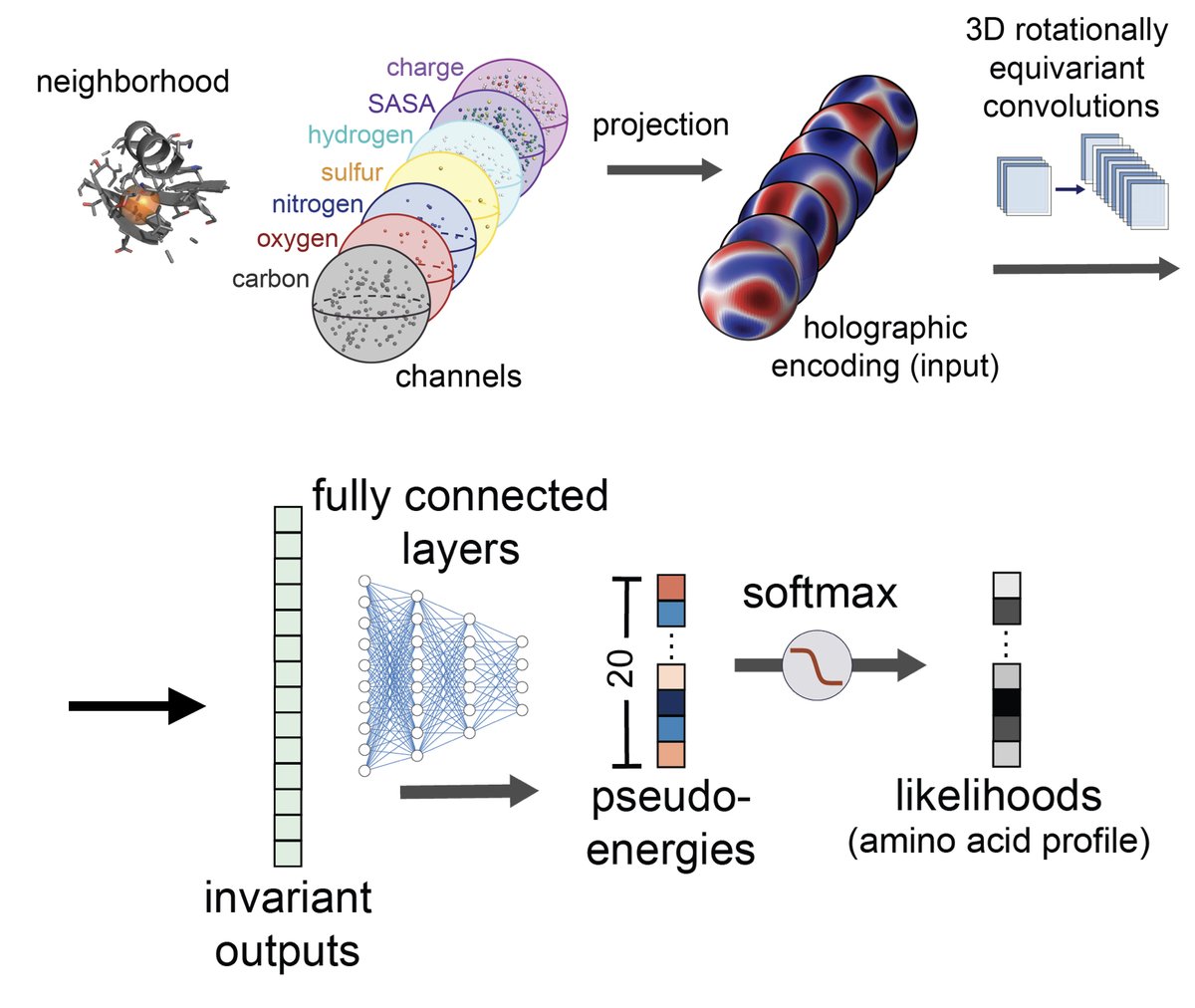
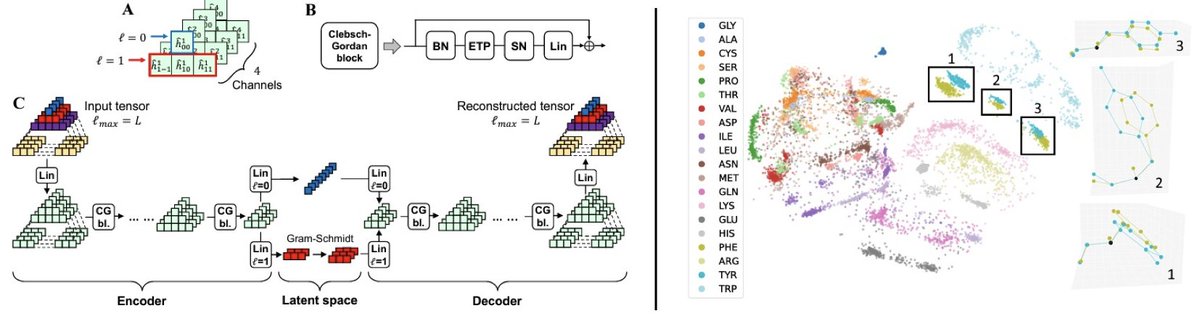
structure-based machine learning model for TCR-pMHC complexes, predicting T-cell affinity to peptide-MHC complexes, quantifying T-cell receptor specificity, and designing de-novo immunogenic peptides:
arxiv.org/abs/2503.00648

English