Tim Grant
32 posts


Tim Grant
@GrantCryoEMLab
cryo-EM structures and methods development, cisTEM development
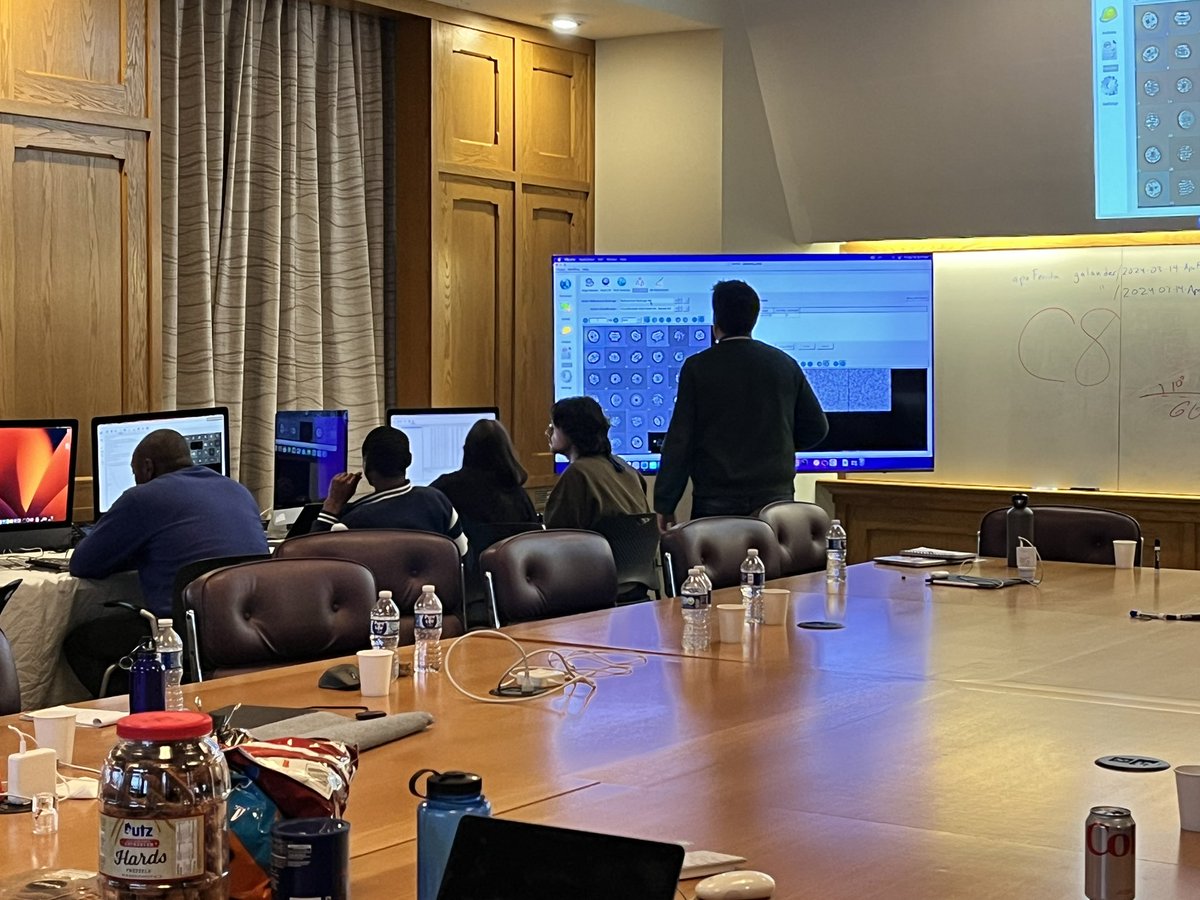







Grad student Peter Ducos was a drone pilot for the US Army. Now he helms the controls for equipment used in #cryoEM research to solve complex problems by exploring molecular structures in exquisite detail. morgridge.org/story/rising-s… #RisingSparks #FearlessScience




Congrats to the IPiB students who received poster awards at the IPiB Retreat! Roma Broadberry (Grant Lab) Jess Davidson (Simcox Lab) Kyle Flickinger (Cantor Lab) Evelyn Okal (Raman Lab) Phoenix Shepherd (Kirchdoerfer Lab) More photos from the day: flic.kr/s/aHBqjAVhZi

Congrats to the IPiB students who received poster awards at the IPiB Retreat! Roma Broadberry (Grant Lab) Jess Davidson (Simcox Lab) Kyle Flickinger (Cantor Lab) Evelyn Okal (Raman Lab) Phoenix Shepherd (Kirchdoerfer Lab) More photos from the day: flic.kr/s/aHBqjAVhZi

















