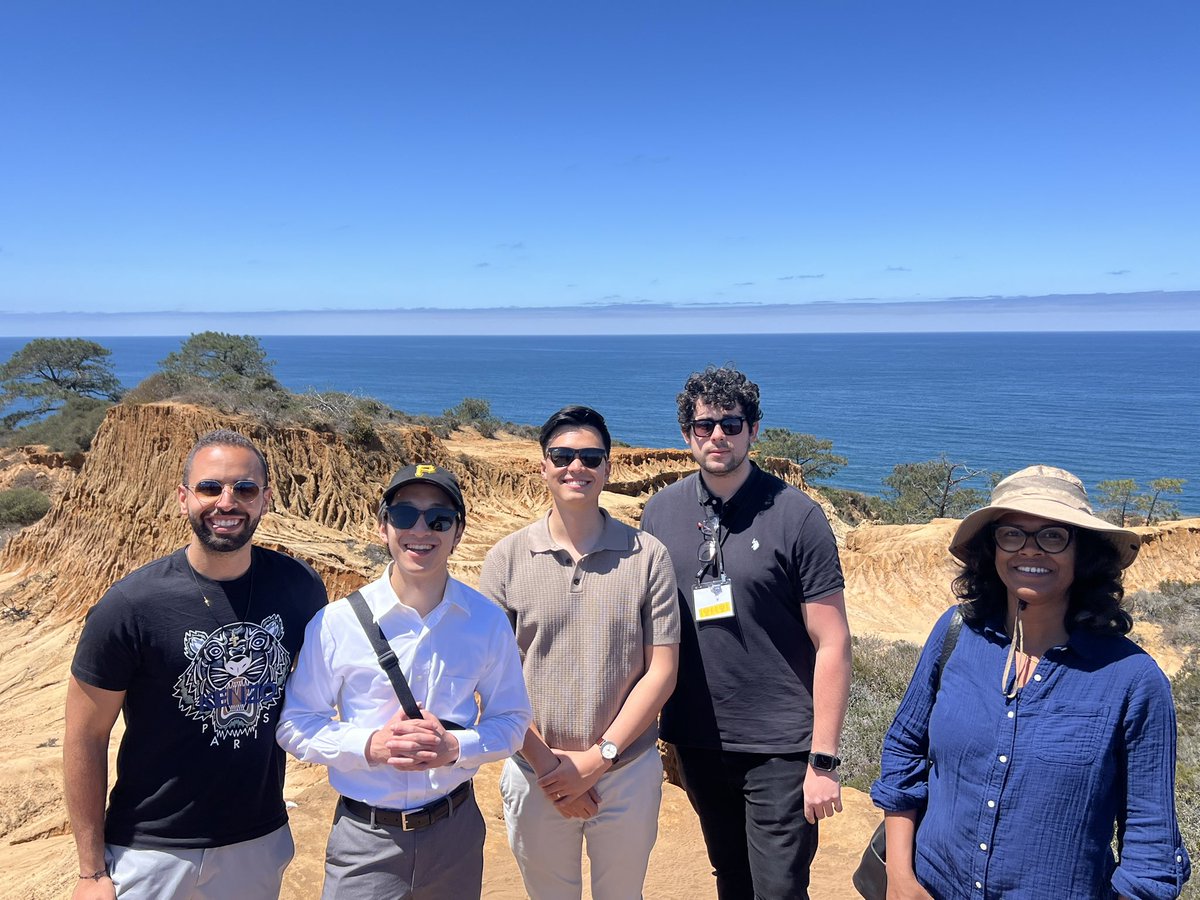
Guglielmo Vetere, MD retweetledi

💣 Important Shift in Cytopenia Grading: CTCAE v6.0 Update
If you are a PI running #Hematology or #Oncology trials, note these key changes in the new CTCAE v6.0 vs v5.0:
📉 General Trend: "Downgrading" of severity. Many counts that were previously Grade 3/4 are now lower grades.
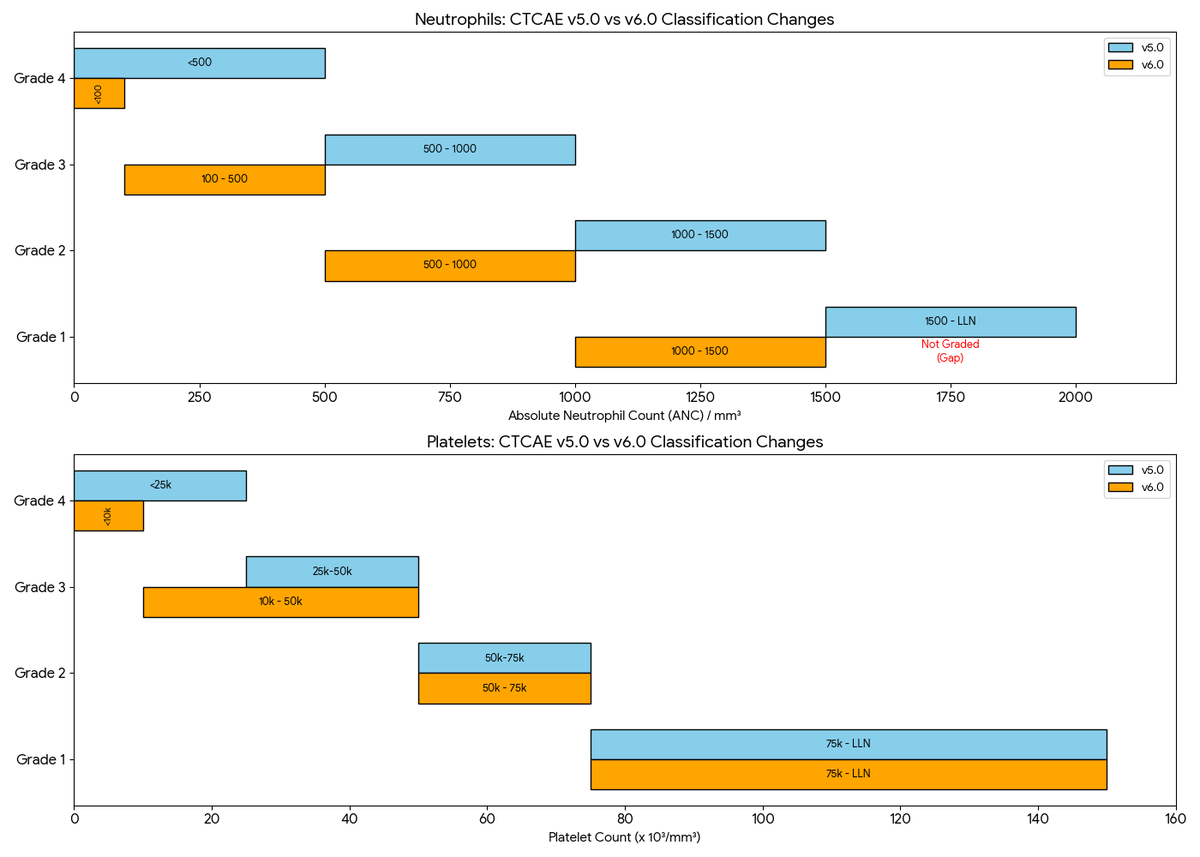
🧪 Neutrophils:
Grade 1 Gone: ANC 1000–1500 is now Grade 1 (was G2). The old Grade 1 (1500–LLN) is no longer graded.
Stricter G4: threshold drops from <500 to <100/mm³.
🩸 Platelets (Thrombocytopenia):
Wider G3: Now covers 10k–50k (previously 25k–50k).
Stricter G4: threshold drops from <25k to <10k/mm³.
v6.0 also adds "transfusion indicated" to Grade 3 and "urgent intervention indicated" to Grade 4 criteria.
Overall, IMO these new thresholds align better with clinical practice.
#ClinicalTrials #MedEd #OncTwitter #DrugSafety
English