Jonathan Funk retweetledi

RFdiffusion3 Now Open Source, Designs DNA Binders and Advanced Enzymes - GEN article
genengnews.com/topics/artific…
English
Jonathan Funk
126 posts


@JonathanFunk12
PhD student - AI for Protein Design Institute for Protein Design


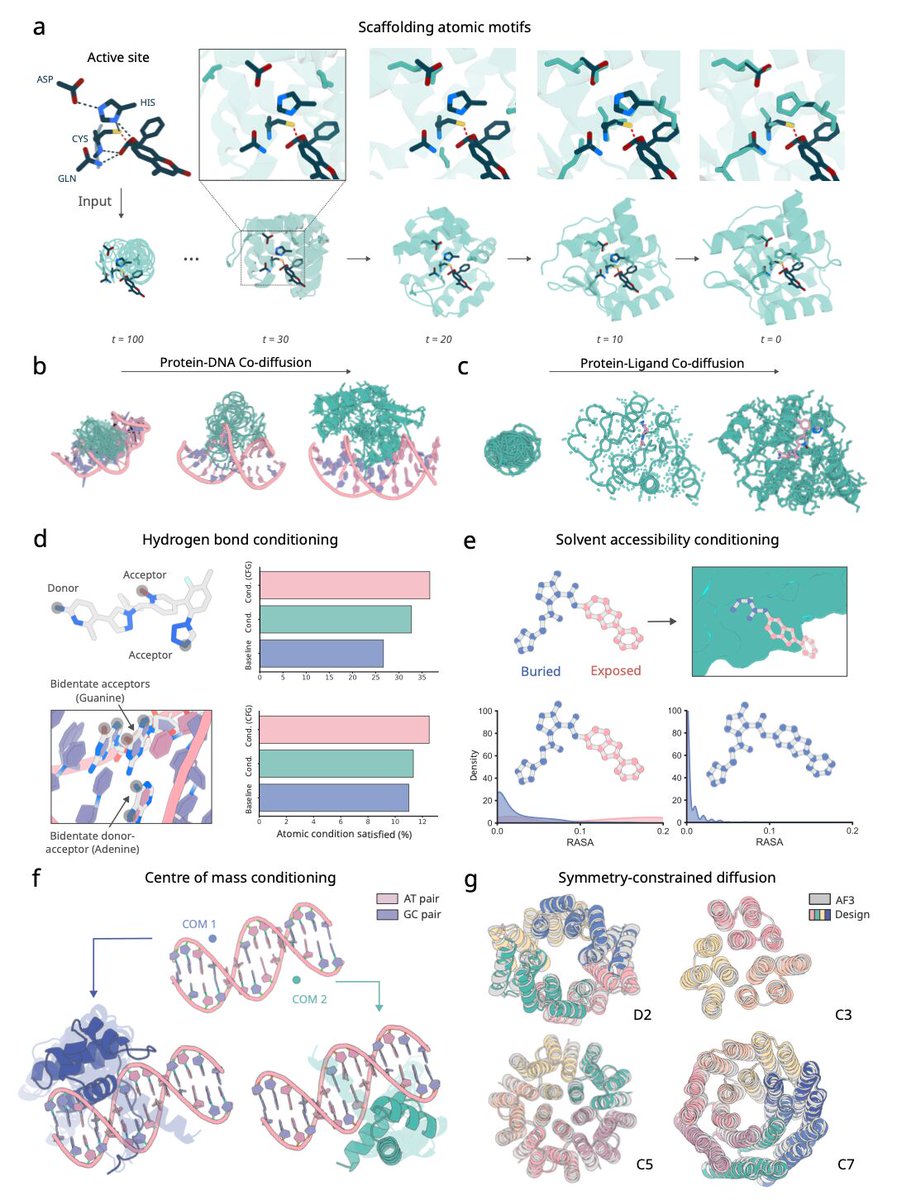



AtomWorks is out! Building upon @biotite_python, we built for a toolkit for all things biomolecules and trained RF3 with it. All open-source, test it via `pip install atomworks`! AtomWorks: github.com/RosettaCommons… RF3: github.com/RosettaCommons… Paper: tinyurl.com/y2w4z65b 1/6








New paper - Transformers, but without normalization layers (1/n)


