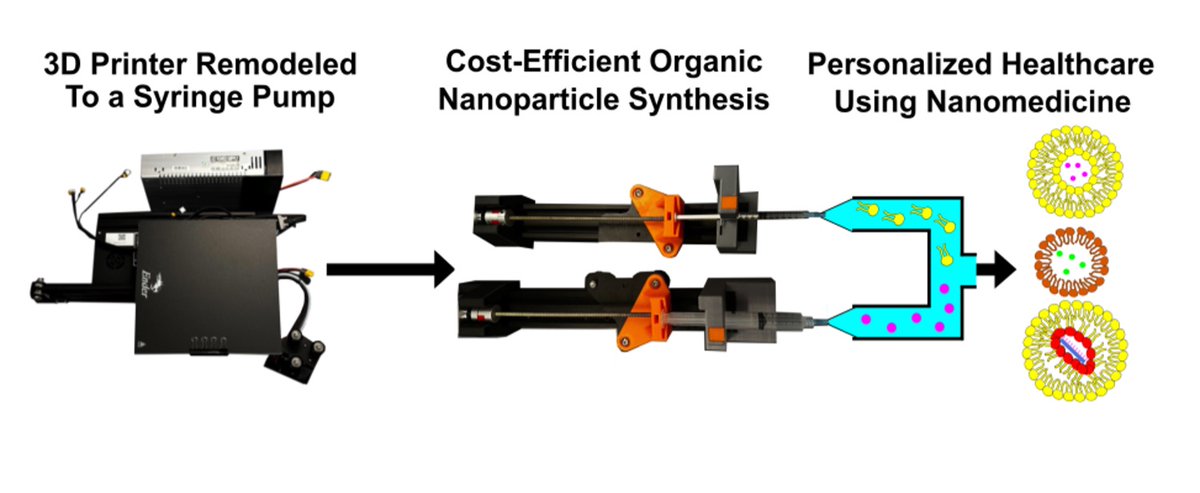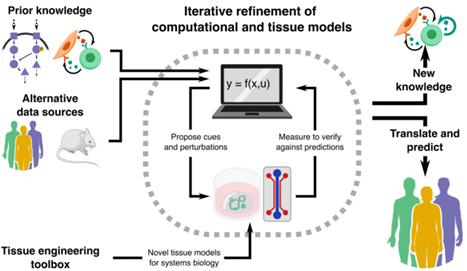
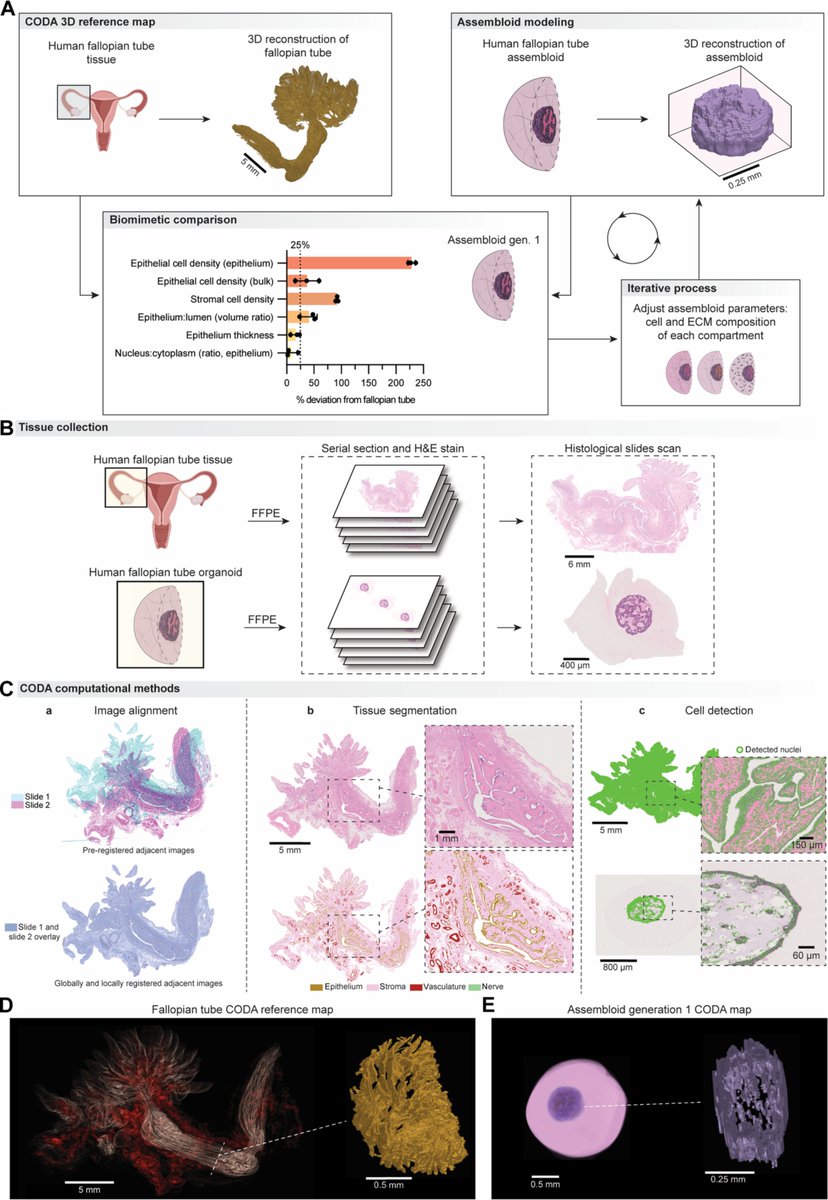
In the pursuit of a better understanding of biological systems and processes, researchers have built increasingly sophisticated physical models to study. #Scilight Learn more 👇 aippub.org/43U9jXO
Jose Cadavid
493 posts

@JoseLCadavid
Chemical engineer messing with systems biology and tissue engineering. Postdoc @MITdeptofBE Lauffenburger/Griffith lab. Dark beer, bass guitar, 3D printing.


In the pursuit of a better understanding of biological systems and processes, researchers have built increasingly sophisticated physical models to study. #Scilight Learn more 👇 aippub.org/43U9jXO









Leveraging Unified Sequence-Structure Representations for Enhanced Protein Stability Prediction 1. A new deep learning framework named ProStab-Former has been introduced to predict protein thermal stability with high accuracy. This model integrates sequence and structure information in a unified representation space, overcoming limitations of previous multi-modal approaches that struggled with indirect information fusion and incomplete capture of sequence-structure interactions. 2. ProStab-Former leverages a pre-trained, multi-modal protein foundation encoder for robust feature extraction at the residue level. It includes Stability-Aware Attention Layers (SAAL) with a structural prior bias and mutation-aware gating, enabling precise capture of local and global stability perturbations induced by mutations. 3. The model incorporates an Epistatic Interaction Module to explicitly model non-additive effects in multi-point mutations. This allows ProStab-Former to efficiently predict the stability changes for numerous single-point mutations in a single pass, significantly accelerating mutation landscape analysis. 4. Experimental evaluations show that ProStab-Former achieves superior performance, with a median Spearman correlation coefficient of 0.84 on the Megascale Test set, surpassing state-of-the-art models like SPURS. It also demonstrates strong generalization across diverse tasks, including melting temperature prediction and pathogenic mutation classification. 5. ProStab-Former's design prioritizes computational efficiency, enabling rapid inference for comprehensive mutation scanning. This makes it highly practical for high-throughput protein engineering and variant effect analysis, positioning it as a valuable tool for both research and industrial applications. 📜Paper: biorxiv.org/content/10.648… #ProteinStability #DeepLearning #ProteinEngineering #Bioinformatics



We are thrilled to share that our first paper from my new lab, Spateo (github.com/aristoteleo/sp…) for spatiotemporal modeling of molecular holograms, is now online in Cell: cell.com/cell/fulltext/…. Spateo is a comprehensive analytical framework for 3D whole-embryo spatiotemporal modeling. Its advanced features include: • 3D alignment and reconstruction at the whole-mouse-embryo scale (see the animation). • 3D spatial domain digitization and cell-cell communication analysis to understand spatial gene expression gradients and both inter- and intracellular communication. • 3D morphometric and volumetric analyses along with 3D morphogenesis vector field modeling to quantify dynamics such as surface area, volume, and cell density across organs, and to dissect the interplay between morphogenesis factors and cell migration. • A “Google Earth”-like browser, Spateo-viewer (viewer.spateo.aristoteleo.com and github.com/aristoteleo/sp…), for interactive and intuitive exploration of 3D spatial data. • Additional features, such as RNA signal-based single-cell segmentation. We are also honored that Nature “News and Views” has highlighted this work as well: nature.com/articles/d4158…. This is really an amazing outcome after two years' heroic revision process that rewrite the entire paper using a new data (doi.org/10.1101/2024.0…) for whole mouse embryos.











In the pursuit of a better understanding of biological systems and processes, researchers have built increasingly sophisticated physical models to study. #Scilight Learn more 👇 aippub.org/43U9jXO