Sabitlenmiş Tweet
Shinsuke Segawa (Maitotoxin)
4.3K posts


Shinsuke Segawa (Maitotoxin)
@MaitotoxicPhD
Chemist, Biologist = Artist. Fluorescent Probe/Click and Bioorthogonal Chemistry Expert. Kyoto U→UT Austin→HKUST→Researcher in Biotech. ケムステスタッフ. 気軽に絡んでね!
Japan→USA→Hong Kong→Japan Katılım Ağustos 2019
1.2K Takip Edilen1.1K Takipçiler
Shinsuke Segawa (Maitotoxin) retweetledi

For readers interested in lipid nanoparticles for oligonucleotide delivery, here's an article covering 60 years of progress in the field
rdcu.be/fb9xD
nature.com/articles/s4157…

English
Shinsuke Segawa (Maitotoxin) retweetledi

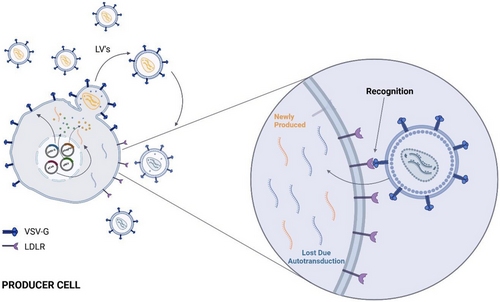
Review: Autotransduction and Pseudotyping as a Key Limitations in Lentiviral Vector Production
…iencejournals.onlinelibrary.wiley.com/doi/10.1002/bi…
Français
Shinsuke Segawa (Maitotoxin) retweetledi

New pre-print from the lab!
Customizing AAVs is usually a slow, trial-and-error process. Leveraging mosaic AAVs, Nolan’s new paper presents MATCH, a "plug-and-play" toolkit that clicks homing proteins directly onto the virus.
>80x better brain transduction
>50% transduction of resting PMBCs
Simple one-pot production
full paper: Receptor-guided AAV Tropism Engineering via MATCH biorxiv.org/content/10.648…

English
Shinsuke Segawa (Maitotoxin) retweetledi

当専攻の大内誠教授が日本化学会第43回学術賞を受賞しました。
題目:側鎖変換による精密高分子のライブラリー合成
chemistry.or.jp/news/informati…
日本語
Shinsuke Segawa (Maitotoxin) retweetledi
Shinsuke Segawa (Maitotoxin) retweetledi

Radical Copolymerization of Formaldehyde with Vinyl Acetate: Direct Incorporation of C–O Bonds in Vinyl Polymer Backbones | Journal of the American Chemical Society pubs.acs.org/doi/10.1021/ja…
English
Shinsuke Segawa (Maitotoxin) retweetledi

Far-red kinase biosensors are applied to image kinase signaling networks at super resolution go.nature.com/3GfqrzL
rdcu.be/faNLM
English
Shinsuke Segawa (Maitotoxin) retweetledi

Entropy-Driven Design of Depolymerizable Polyolefins from Strained Bridged Bicyclic Monomers | Journal of the American Chemical Society pubs.acs.org/doi/10.1021/ja…
English
Shinsuke Segawa (Maitotoxin) retweetledi

DNA–drug conjugates enable logic-gated drug delivery amplified by hybridization chain reactions go.nature.com/4bVNs6Z
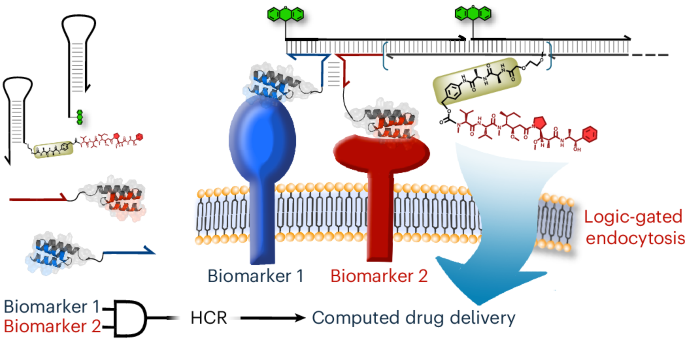
English
Shinsuke Segawa (Maitotoxin) retweetledi

This paper took us 13 years and is one of the longest papers ever in Cell. Check it out & judge for yourself
cell.com/cell/fulltext/…

Olafur Pall Olafsson@olafurpall80
@JoinLifespan Just because epigenetic drift follows a predictable pattern doesn't necessarily mean it's causal in aging. Predictable patterns can form from stochastic reactions. Also you cannot fix all extracellular damages with cellular rejuvenation. More here: olafurpall.substack.com/p/why-aging-is…
English
Shinsuke Segawa (Maitotoxin) retweetledi

Have you heard of “zombie cells”?
@Nature just published a commentary that will make you rethink what "life" means.
Scientists from JCVI took bacterial cells, killed their genomes with a chemotherapy drug (mitomycin C crosslinks the DNA — permanently disabled), then transplanted a synthetic genome from a completely different species into these dead cell bodies.
Some rebooted. Grew. Divided. Alive again — but running entirely on foreign DNA.
The cell body is hardware. The genome is software. They wiped the drive, installed a new OS, and the machine booted.
What makes this technically distinct from prior JCVI work: previous genome transplants competed with the host's native genome, requiring antibiotic selection to find successful transfers. Here, the host genome is already dead — only cells revived by the new genome survive. No selection markers, no competition. Cleaner, more scalable.
The historical arc is worth appreciating:
• 2007: genome transplant between species
• 2010: first synthetic self-replicating cell
• 2016: minimal genome (473 genes — the floor of life)
• 2026: dead chassis + synthetic genome = living cell
The question this raises isn't just technical. A M. capricolum body running a M. mycoides genome — what species is it? The answer is neither, which means species identity may live in the genome, not the cell.
That has implications far beyond bacteria.
Paper link: nature.com/articles/d4158…

English
Shinsuke Segawa (Maitotoxin) retweetledi

遺伝子変異に合わせたRNA標的創薬へ RNA構造と低分子化合物の相互作用を大規模に検出する新技術 『BIVID-MaP』を開発 kyoto-u.ac.jp/ja/research-ne…
日本語
Shinsuke Segawa (Maitotoxin) retweetledi

CRISPR–Cas9 might offer a way to make safer, more effective cancer-fighting immune cells engineered inside the human body
go.nature.com/4rMD8nI
English
Shinsuke Segawa (Maitotoxin) retweetledi

CART 細胞による腫瘍免疫療法において、CART細胞の代謝を活性化し、結果として腫瘍免疫を増強する物質として3-ヒドロキシ酪酸を同定したという論文。
cell.com/cell/fulltext/…
日本語
Shinsuke Segawa (Maitotoxin) retweetledi

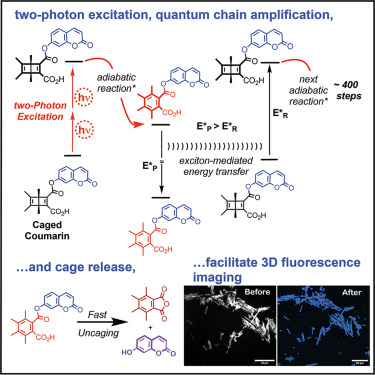
Online now: Three-dimensional imaging via quantum chain amplification in a crystalline two-photon adiabatic photocage dlvr.it/TRcFPB
English
Shinsuke Segawa (Maitotoxin) retweetledi

170種類にものぼるRNA修飾とその生物学的意義についての総説。化学構造とかも書いてあるので、私のような門外漢には助かる。CELL誌。
cell.com/cell/fulltext/…
日本語
Shinsuke Segawa (Maitotoxin) retweetledi

📄 Nature Communications に論文が公開されました!
BIVID-MaP: 1塩基の遺伝子変異が、RNA標的低分子医薬の結合をどう変えるかを世界で初めて体系的に検出できる技術を開発しました。
xFOREST社、京大CiRA/東大齊藤研、東北大永次研との共同研究成果です。
🔗 nature.com/articles/s4146…
日本語
Shinsuke Segawa (Maitotoxin) retweetledi

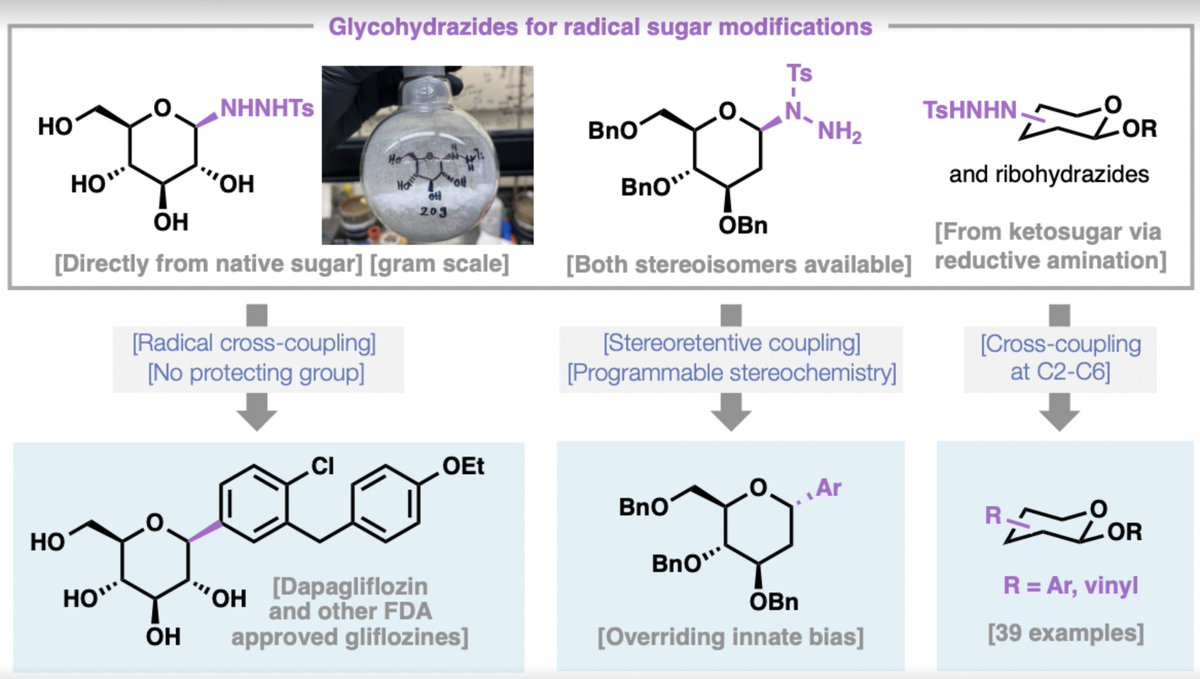
Making C-glycosides SWEET and simple! Today in @ChemRxiv we disclose (chemrxiv.org/doi/full/10.26…), in collaboration with @GroupAggarwal, an incredibly easy way to achieve radical functionalization of sugars. In this video (youtu.be/Fqdbgmx7zEI), a two-step synthesis of the billion dollar drug Dapagliflozin is achieved using household vinegar and dextrose powder from the local supplement store.
High Level Summary:
The work addresses a longstanding challenge in carbohydrate chemistry: the efficient, scalable, and stereocontrolled synthesis of C-aryl glycosides directly from unprotected native sugars. C-Aryl glycosides form the core pharmacophore of the SGLT2 inhibitors (dapagliflozin, canagliflozin, empagliflozin, and related agents), which are frontline therapies for type 2 diabetes and represent one of the highest-grossing classes of small-molecule drugs.
Conventional synthetic routes to these molecules generally require extensive protecting-group manipulations, multi-step activation of glycosyl donors, or organometallic additions under demanding conditions. Recent advances in radical and transition-metal-catalyzed cross-couplings have improved access, yet most approaches still depend on protected precursors, specialized reagents, or protocols that are difficult to scale.
We report a practical alternative based on glycosyl sulfonyl hydrazides—stable, crystalline radical precursors that are prepared in a single step from unprotected sugars by treatment with tosylhydrazine in acetic acid, followed by simple crystallization. These hydrazides undergo redox-neutral nickel-catalyzed radical cross-coupling with aryl iodides or bromides under mild conditions (70 °C, DMSO, tetramethylguanidine as base). The reaction requires no external oxidant or reductant, no photocatalyst, and no organotin species. In glucose-derived systems the coupling typically delivers high β-selectivity (>19:1 in many cases), an outcome that appears to depend on hydrogen-bonding interactions between tetramethylguanidine and the free hydroxyl groups.
The main findings are as follows:
All five FDA-approved SGLT2 inhibitors, as well as several clinical candidates, can be prepared in a single coupling step from the corresponding glycohydrazide.
Decagram-scale synthesis of dapagliflozin was demonstrated starting from commercial dextrose; the product was isolated by aqueous workup and recrystallization (no column chromatography required at this scale).
Di- and trisaccharides (lactose, cellobiose, maltose, maltotriose) couple directly to give aryl-linked oligosaccharides.
Several natural products and medicinally relevant structures (salmochelin-SX, neopetrosin C, the tryptophan-mannose conjugate, and a ribose-derived IMPDH inhibitor) that previously required 9–20 steps or costly reagents are now accessible in 1–4 steps with good stereocontrol.
The platform extends to non-anomeric C–C bond formation at positions C2–C6 on glucose and ribose scaffolds, providing the first systematic exploration of radical diversification across these positions.
Stereoretentive radical cross-coupling, using configurationally pure hydrazides, enables programmable delivery of either α- or β-anomers, overriding inherent substrate biases and providing access to stereoisomers not previously obtainable by radical methods.
The chemistry builds on our earlier development of sulfonyl hydrazide-based redox-neutral cross-coupling and stereoretentive radical arylation, here adapted and optimized for carbohydrate substrates. The method is operationally straightforward, uses inexpensive reagents and starting materials, and eliminates protecting-group strategies.

YouTube
English