Samuel G. Huete
1.7K posts


Samuel G. Huete
@MicroBioMol
🧪 Molecular microbiologist 🧫 at @einlabryc ☣️ Infectious diseases and 🧬 evolution 🔬 PhD from @InstitutPasteur 🌐 Head of @JISEM_SEM 🎶 Composer & ⛰️hiker!



😱😱¿Ciencia ficción o realidad? 🧪🧟♂️ ¡Científicos han logrado "revivir" bacterias muertas 🦠✝️! Un nuevo estudio del J. Craig Venter Institute marca un hito en la biología sintética. No es solo edición genética, es "reanimación" celular. 🧵 ↘️ 1️⃣ Células "Zombie": Los investigadores mataron bacterias (M. capricolum) desactivando su ADN con químicos, dejando solo la estructura celular intacta 🧫. 2️⃣ Trasplante Total: Insertaron un genoma sintético completo en estas bacterias muertas 🧬. 3️⃣ El Despertar: Al recibir el nuevo ADN, la bacteria "muerta" volvió a la vida, empezó a crecer y a dividirse como una especie nueva ⚡. 4️⃣ Sin Antibióticos: A diferencia de métodos antiguos, este sistema no necesita marcadores de selección, lo que lo hace mucho más eficiente para crear organismos a la carta 🛠️. Esto abre la puerta, por ejemplo, a crear microbios diseñados para producir combustibles limpios o medicamentos específicos. 🌍💊 🔗 @SEMicrobiologia @COSCEorg @ANIH_1 @microBIOblog #BiologíaSintética #Ciencia #Genética #Innovación #CraigVenter #Biotecnología #Microbiología 👇 biorxiv.org/content/10.648…

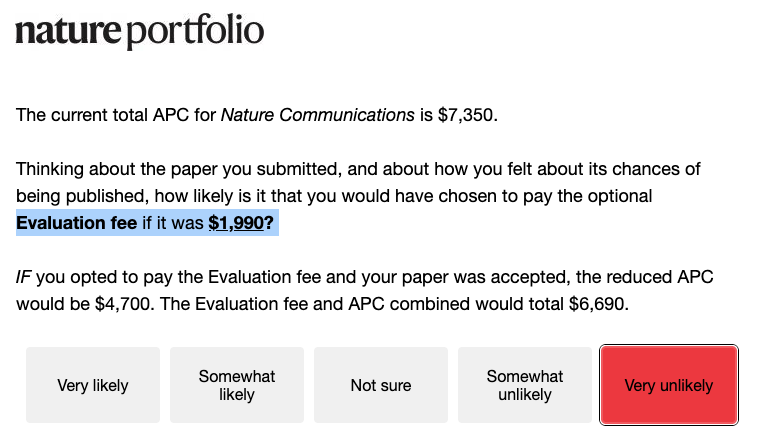





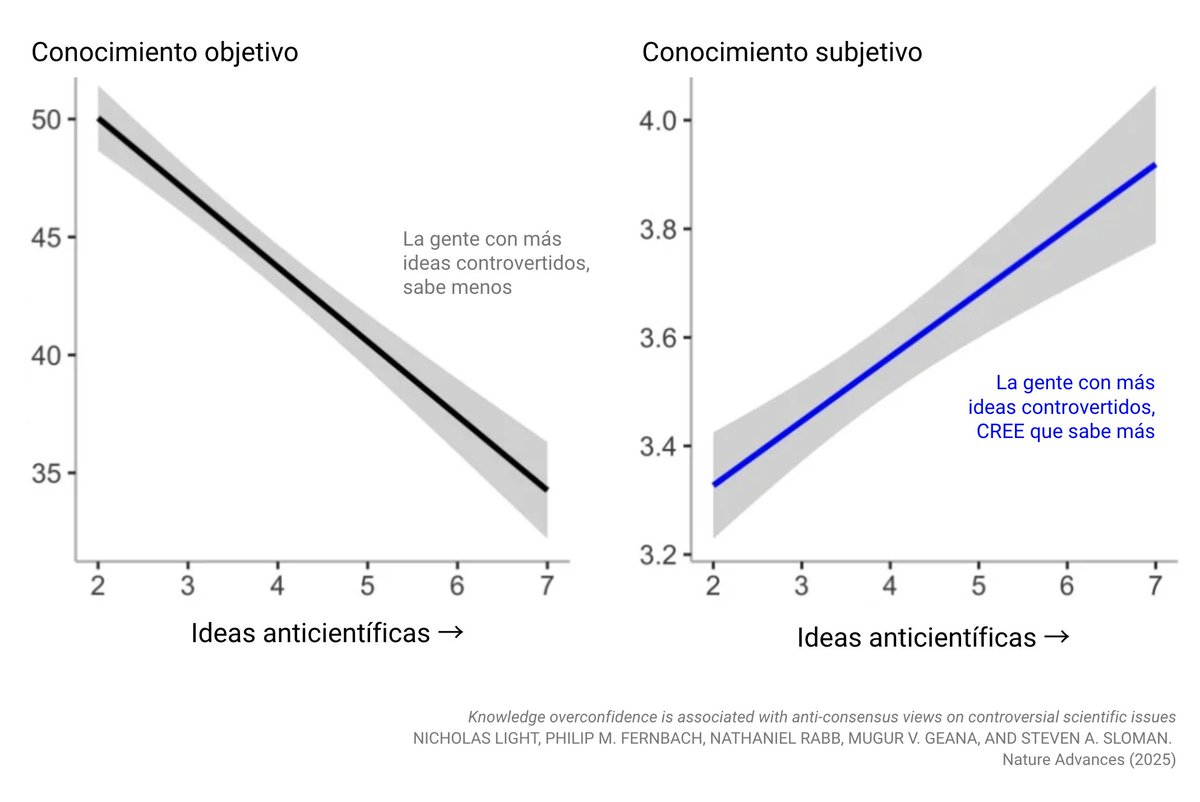
For decades, peer review has been treated as the gold standard of scientific validation. Yet many scientists know the reality: the system is far from perfect. Peer review is broken and sometimes even corrupted. The process can be slow, inconsistent, and vulnerable to bias. Reviewers are sometimes asked to judge work outside their true expertise. In other cases, they may be evaluating ideas that challenge the very paradigm in which they were trained. And occasionally, reviewers are simply competitors. Ironically, the most prestigious journals can also be the most conservative. Truly new ideas are often met with skepticism, while safer work that fits the current narrative moves more easily through the system. Increasingly, papers are judged less by the originality of the idea and more by the volume of data, the sophistication of statistics, and the beauty of the figures. Science risks becoming data-rich but idea-poor. But there is an important reality to remember: journals do not ultimately decide the impact of scientific work. Impact is decided later, by the community. By the scientists who read it, test it, debate it, and cite it. In the end, citations and ideas determine the legacy of a paper, not the impact factor of the journal that first published it. Science has always advanced by questioning assumptions. Perhaps it is time we also question the system that filters scientific ideas.

I'm currently writing a scientific grant and sometimes wonder how much effort we invest making our research ideas "attractive" to reviewers instead of focusing on having good ideas. I wish we could get funded after a good and calm conversation with an evaluator over some coffee


📢 ¡Atención! 🔬 Recordad que está abierta la convocatoria del Programa de Ayudas de Movilidad César Nombela 2026 de la @SEMicrobiologia 👩🔬👨🔬 Una gran oportunidad para estancias cortas para pre y postdocs🦠 - Deadline: 8 de Marzo!!! 🔗 Toda la info aquí: semicrobiologia.org/becas/programa…




"Los números especiales de MDPI y Frontiers son un enorme coladero. La ANECA debe excluir explícitamente estos trabajos. Universidades y OPI deben identificar las carreras construidas mediante números especiales y penalizarlas”, afirma @isidroaguillo elpais.com/ciencia/2026-0…