Aanchal
1.4K posts


Aanchal
@Origichals
Human | Learner | PhD researcher in PTMs involved in DNA repair using AlphaFold and structural biology approach @MSuskiewicz lab @CNRS.

This paper is wild. After 3 rounds of directed evolution, they converted a DNA polymerase into an enzyme that can do: - RNA synthesis - Reverse transcription - Synthesis of "unnatural" nucleotides - Synthesis of DNA-RNA chimeras One of the best papers I’ve read recently. For context: In nature, it is DNA polymerase that takes a DNA sequence as a template and then copies it. These enzymes are crucial in replicating the genome for cell division, and they are EXTREMELY specific for DNA over RNA. This is key because RNA nucleotides are present in the cell at concentrations ~100x higher than DNA nucleotides, so the enzyme has evolved clever strategies to select one over the other. RNA polymerases, for comparison, are the enzymes that take a DNA sequence as template and then convert it into RNA. They are involved in gene expression, for example. To convert a DNA polymerase into an RNA polymerase (and all the other functions I mentioned earlier), the authors did a fairly straightforward directed evolution experiment. First, they took four DNA polymerase enzymes belonging to various archaea. These DNA polymerases don’t check for DNA vs. RNA as stringently as other types of cells, so they’re a good starting point to evolve RNA polymerases. The authors inserted some targeted mutations into these enzymes, based on known mutations in the literature. For example, they swapped the amino acid at position 409 for a smaller amino acid, thus removing a “gate” that keeps RNA building blocks from entering the enzyme. Next, they took the four genes encoding these DNA polymerases and cut them up into 12 segments each. They randomly stitched these 12 segments together — from the four different genes — to build millions of unique variants. Each shuffled gene was inserted into an E. coli cell. Then, they grew up these cells (each carrying a unique polymerase) and put them into microfluidic droplets. A device isolates each droplet, lyses the cell open, and releases the polymerase. The droplet also contains RNA building blocks and a DNA template, encoding a fluorescent reporter. If the polymerase begins synthesizing RNA, it will produce a detectable signal. They screened about 100 million droplets in 10 hours of work, searching for those with a signal. For each well that yields a fluorescent signal, the researchers isolated the DNA and sequenced it to figure out which polymerase it was. They repeated this 3x times, finally isolating a really excellent RNA polymerase variant which they called "C28." C28 has 39 mutations compared to the wildtype enzymes. It incorporates about 3.3 nucleotides of RNA per second, with 99.8% fidelity. The crazy thing is that this enzyme can also copy DNA or RNA templates back into DNA (reverse transcription), or use chimeric DNA-RNA molecules as a template and amplify them. It is just a super versatile polymerase that can act on DNA, RNA, or modified nucleotides, to build just about anything.

This paper is wild. After 3 rounds of directed evolution, they converted a DNA polymerase into an enzyme that can do: - RNA synthesis - Reverse transcription - Synthesis of "unnatural" nucleotides - Synthesis of DNA-RNA chimeras One of the best papers I’ve read recently. For context: In nature, it is DNA polymerase that takes a DNA sequence as a template and then copies it. These enzymes are crucial in replicating the genome for cell division, and they are EXTREMELY specific for DNA over RNA. This is key because RNA nucleotides are present in the cell at concentrations ~100x higher than DNA nucleotides, so the enzyme has evolved clever strategies to select one over the other. RNA polymerases, for comparison, are the enzymes that take a DNA sequence as template and then convert it into RNA. They are involved in gene expression, for example. To convert a DNA polymerase into an RNA polymerase (and all the other functions I mentioned earlier), the authors did a fairly straightforward directed evolution experiment. First, they took four DNA polymerase enzymes belonging to various archaea. These DNA polymerases don’t check for DNA vs. RNA as stringently as other types of cells, so they’re a good starting point to evolve RNA polymerases. The authors inserted some targeted mutations into these enzymes, based on known mutations in the literature. For example, they swapped the amino acid at position 409 for a smaller amino acid, thus removing a “gate” that keeps RNA building blocks from entering the enzyme. Next, they took the four genes encoding these DNA polymerases and cut them up into 12 segments each. They randomly stitched these 12 segments together — from the four different genes — to build millions of unique variants. Each shuffled gene was inserted into an E. coli cell. Then, they grew up these cells (each carrying a unique polymerase) and put them into microfluidic droplets. A device isolates each droplet, lyses the cell open, and releases the polymerase. The droplet also contains RNA building blocks and a DNA template, encoding a fluorescent reporter. If the polymerase begins synthesizing RNA, it will produce a detectable signal. They screened about 100 million droplets in 10 hours of work, searching for those with a signal. For each well that yields a fluorescent signal, the researchers isolated the DNA and sequenced it to figure out which polymerase it was. They repeated this 3x times, finally isolating a really excellent RNA polymerase variant which they called "C28." C28 has 39 mutations compared to the wildtype enzymes. It incorporates about 3.3 nucleotides of RNA per second, with 99.8% fidelity. The crazy thing is that this enzyme can also copy DNA or RNA templates back into DNA (reverse transcription), or use chimeric DNA-RNA molecules as a template and amplify them. It is just a super versatile polymerase that can act on DNA, RNA, or modified nucleotides, to build just about anything.




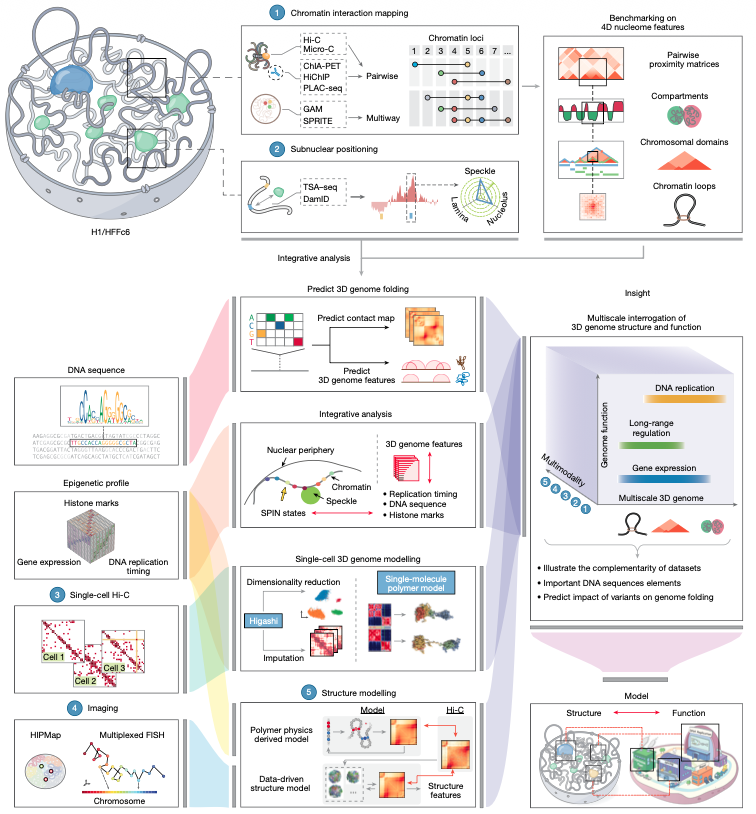



Protein language models just got an upgrade. Meet Profluent-E1: a free, flexible, frontier protein sequence encoder. E1 is built with retrieval augmentation to learn from multiple sequences. Models trained over 4T tokens with only 150M-600M params, E1 is SOTA for zero-shot functional and unsupervised structural tasks. It raises the bar for protein representation learning and is freely available today.





Favourite examples where pi appears unexpectedly?

Happy to share our study showing that the Ub E3 ligase SIAH1, known to dimerise via its C-term SBD, also dimerises via its N-term RING. When these tendencies combine, fl SIAH1 forms multimers, which might explain its clustering in cells & preference for multimeric substrates.



The life of each one of us began when a sperm and an egg came together. But what actually happens at a molecular level? Our latest work @PauliGroup @CellPress reveals a conserved fertilization complex that bridges sperm and egg in vertebrates! (1/8) cell.com/cell/fulltext/…