
OxASN: The Oxford Antigen Specificity Network
362 posts


OxASN: The Oxford Antigen Specificity Network
@Ox_T_Cell_ASN
OxASN is a multidisciplinary network of scientists with the common gaol of decoding the principles of antigen specific cellular immunity in time and space.


If this would be helpful to anyone, but you don't want to buy a copy, you can read it for free online: Link: heyzine.com/flip-book/a5dd… Password: BeKind

Our new book "Self-Doubt" is written by scientists for scientists. You might think you are alone when you doubt your ability or suitability for science - you aren't. We all question ourselves and carry doubt with us daily. amazon.com/Self-Doubt-Ant…




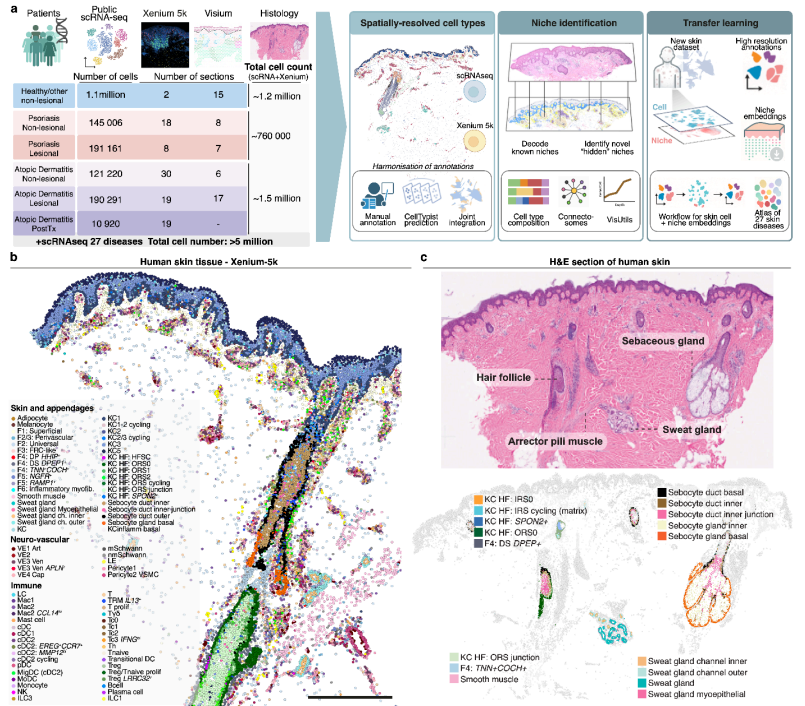









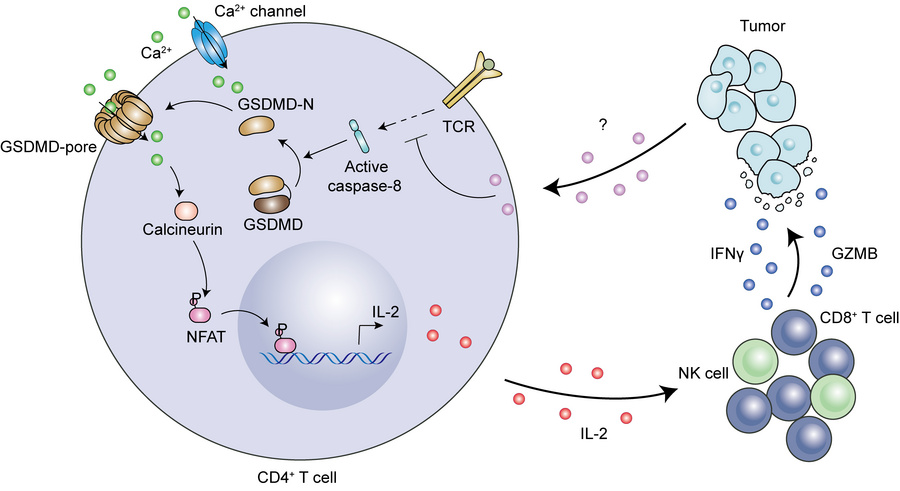




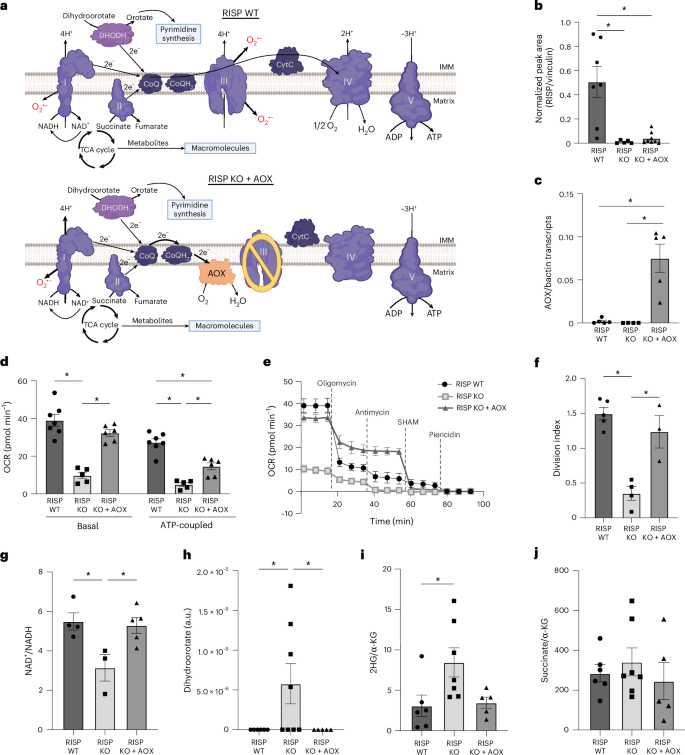



ABSTRACT submission deadline now extended to July 11th! Register now to join us at this exciting Meeting and Hackathon. cvent.me/94L9A5 Oxford, Sept 29th - Oct 1st 2025 #Tcells #AI #Immunology #Oxford #Hackathon