Sabitlenmiş Tweet

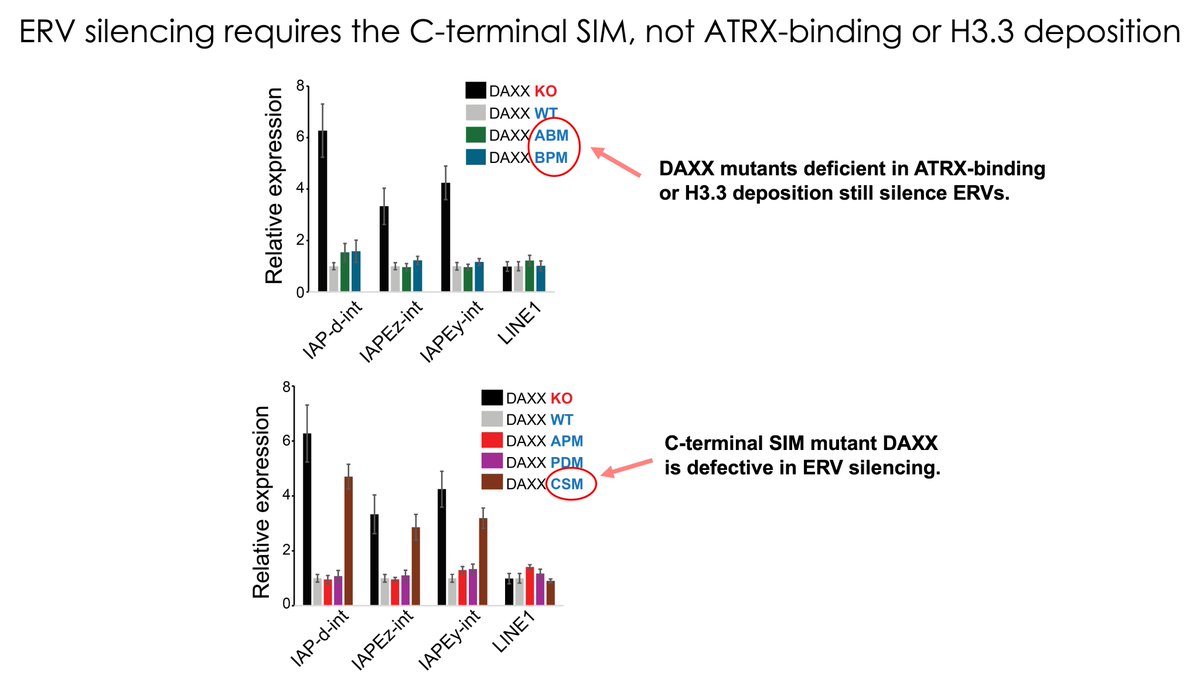
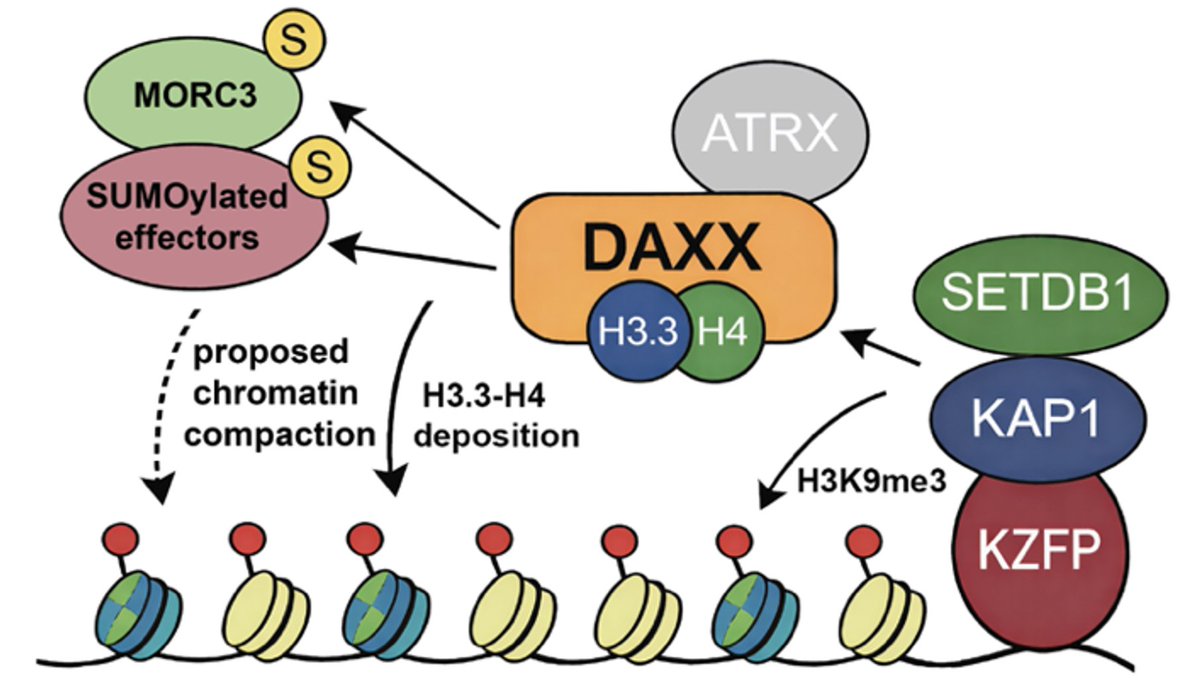
New preprint from our lab! An H3.3 knockout does two things at once: it removes H3.3 from chromatin and destabilizes DAXX. We disentangle those functions and find that DAXX-mediated H3.3 deposition can be uncoupled from ERV silencing.
biorxiv.org/content/10.648…
English