Luis I. Gutiérrez-Rus
271 posts


@RusLuis
Postdoctoral Research Associate | NNF Center for Protein Design @koebenhavns_uni | Protein & Enzyme Design • Evolution • Engineering








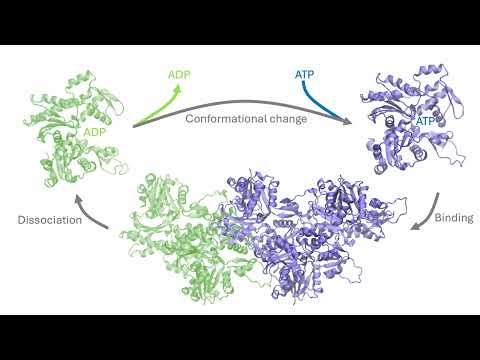
Today in the journal Science: BioEmu from Microsoft Research AI for Science. This generative deep learning method emulates protein equilibrium ensembles – key for understanding protein function at scale. msft.it/6010S7T8n

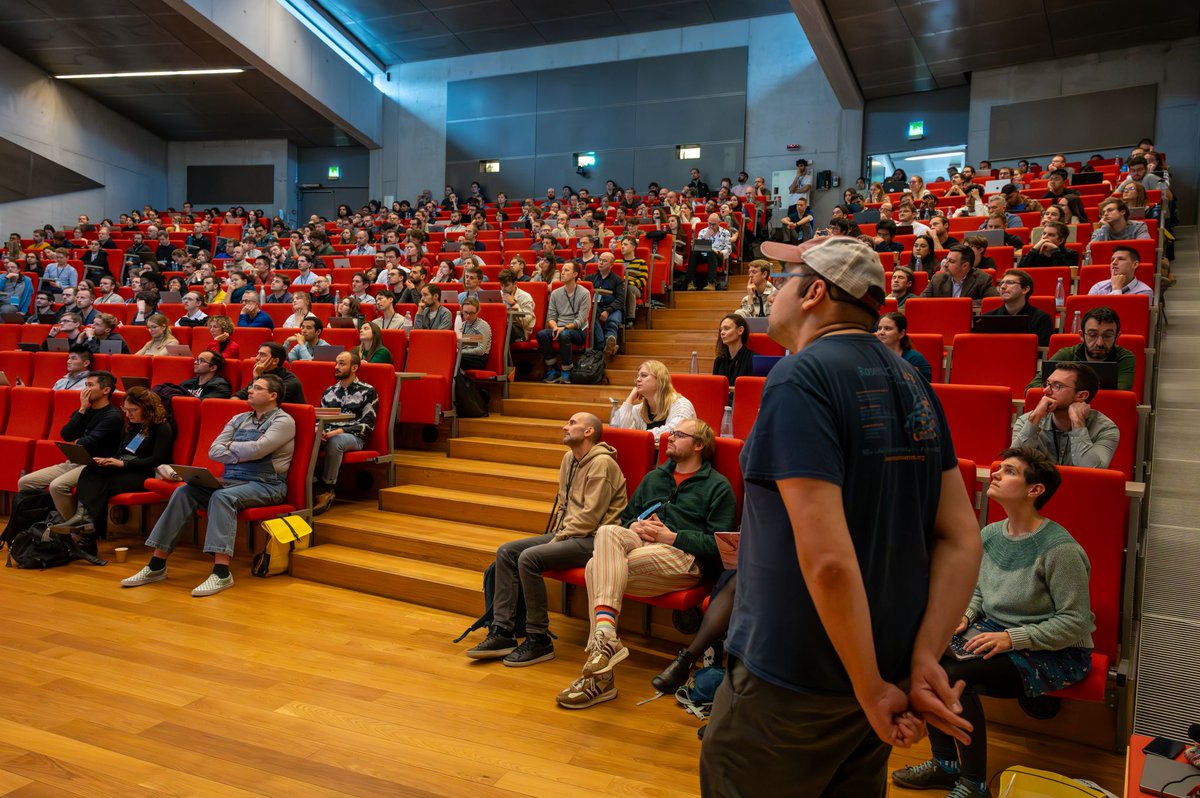
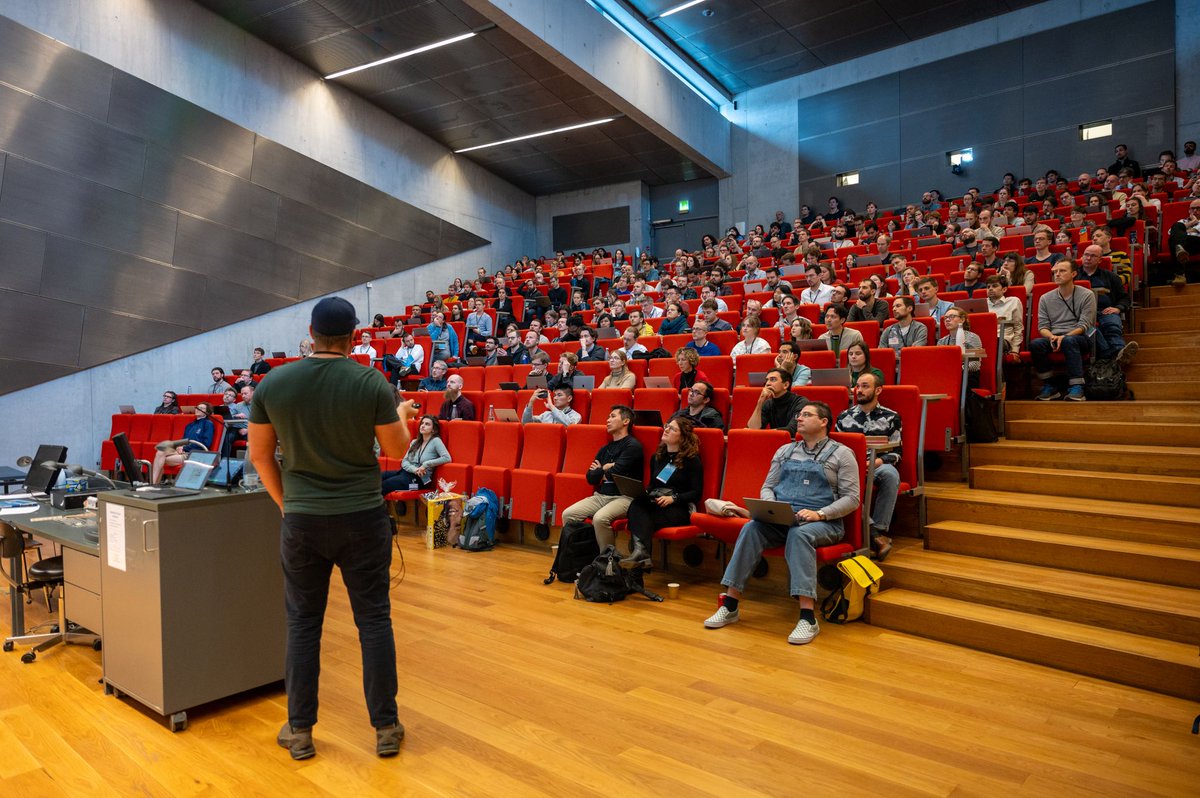
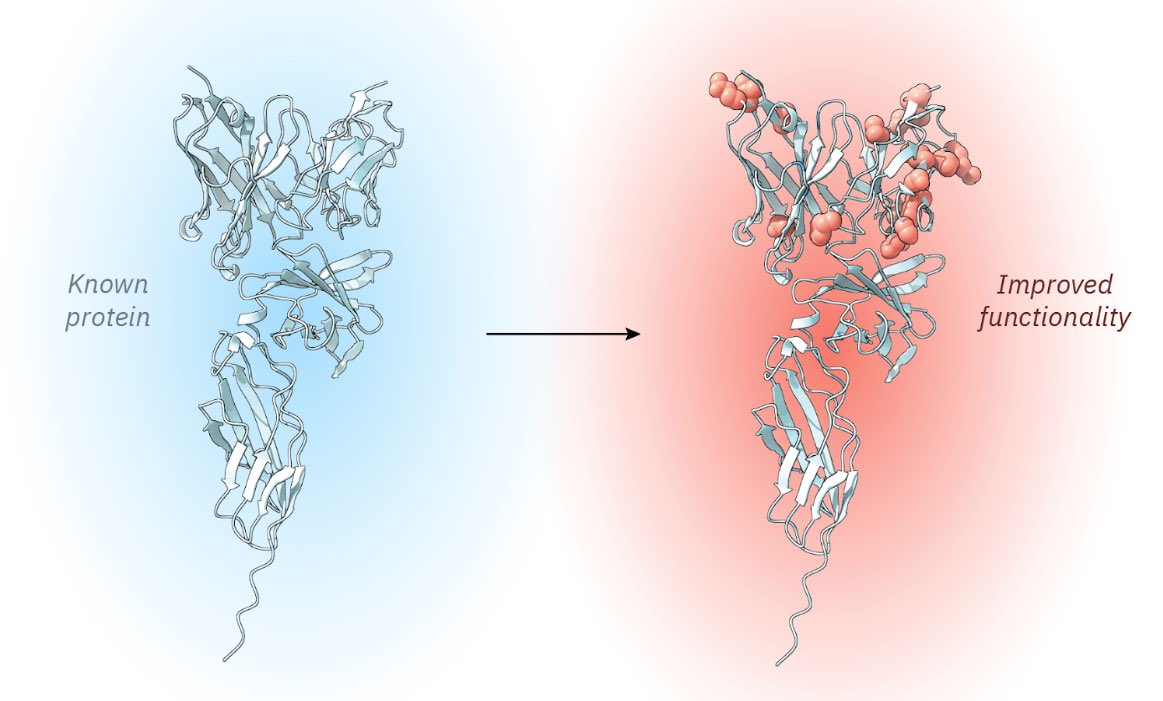
Designing proteins from scratch opens exciting new possibilities for solving global challenges in health and sustainability. That is why we are funding an ambitious new Center for Protein Design at @koebenhavns_uni. Led by Professor Dek Woolfson, world-renowned in this field, the centre aims to build on recent breakthroughs to establish a powerhouse for protein design in Europe. Read more: novonordiskfonden.dk/en/news/new-da…