Sabitlenmiş Tweet

Paper @PNASNews
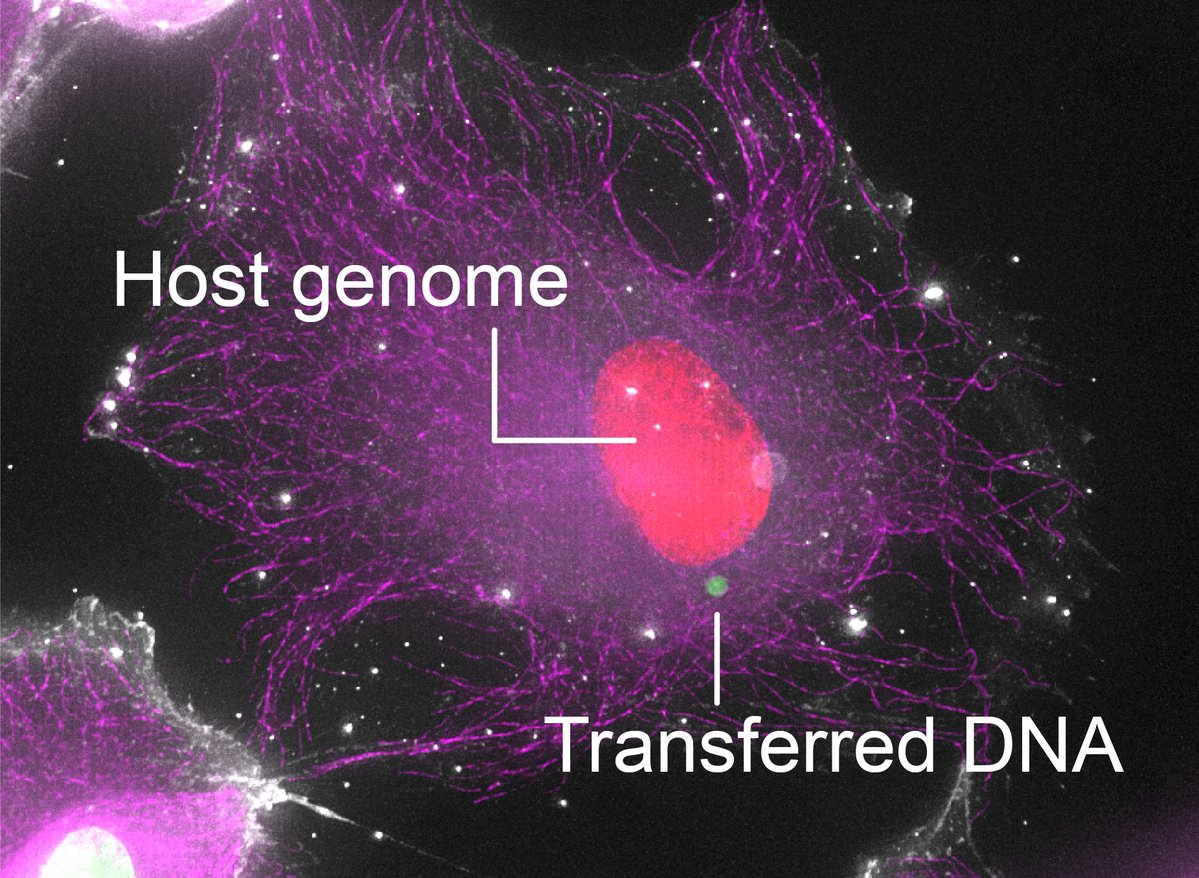
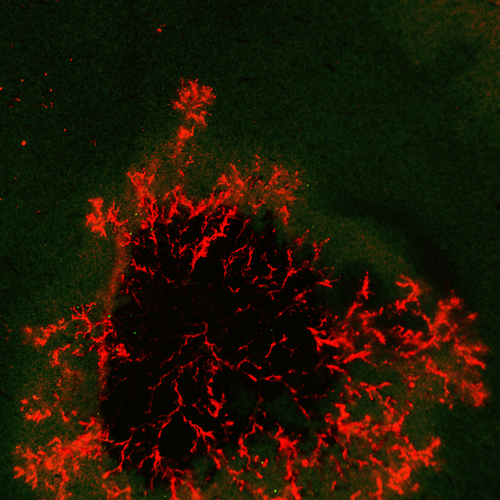
We think nematodes break open plant cells, sense small molecules (termed effectostimulins), which switch on a master regulator SUGR1, which switches on effectors, which break cells.
Looks like a feed-forward loop driving infection.
pnas.org/doi/10.1073/pn…
English