
Smell is the least-understood of our senses. With @jdthamores and @SeyoneC, we trained biologically-inspired models that first map odorant molecules to their receptor activation profiles and subsequently predicts their odor percepts.
Seyone Chithrananda
2.1K posts
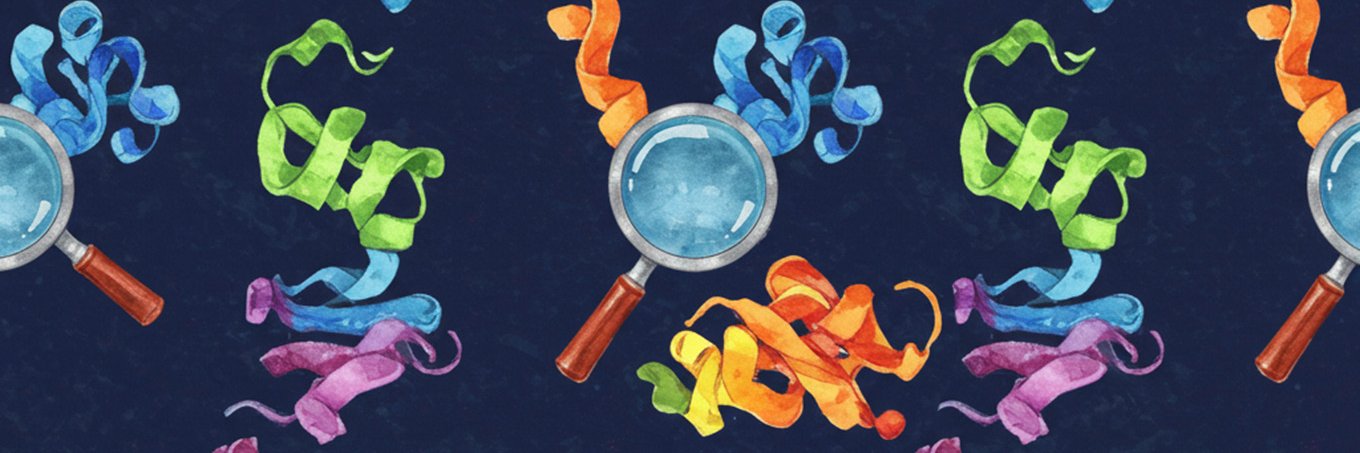


@SeyoneC
🇨🇦 | 1st yr phd @stanford, bear @ucberkeley

Smell is the least-understood of our senses. With @jdthamores and @SeyoneC, we trained biologically-inspired models that first map odorant molecules to their receptor activation profiles and subsequently predicts their odor percepts.



The Liberal party has Patrick Pichette a former Senior VP of Google on stage who lives in Europe by the way, say that if Canadians want to leave Canada to work in the US they need to pay an exit tax of half a million dollars. The guy did the very thing to get a Microsoft job decades ago and paid 30 bucks. Now he wants young Canadians to be trapped here. The Liberals are nuts.



how is this a class? absolutely insane line-up

"Why does our FDA still incentivize all of this innovation to go to China?" @zachweinberg: "You can go to China, you can run a first-in-human study in a Chinese population at a Chinese hospital, you get your result, and then you can take that result back to America and skip the line." "I don't have to redo that Phase 1 and Phase 2 in a Western nation. I can use my Chinese data to open a Phase 3 study here and go for an approval." "Think about the incentive structure for a US biotech. You have to go to China. There is no alternative path because you've got competition on the other side who is racing ahead with infrastructure that you can't use." "We don't inspect, we don't audit, we don't send inspectors to these clinical trial sites. We have no idea what's actually going on." The Hill & Valley Forum 2026 @HillValleyForum






“One of the great things about Iso is how interdisciplinary everyone is, people come from so many different backgrounds.” We spoke with Bilal Khan, Software Engineer at @IsomorphicLabs, about our unique, collaborative culture and frontier AI built to tackle the world's most complex problems. Watch the video and head to the comments to explore our current roles.

Our lying Ontario premier has just stolen $50 from every single person in Ontario. The estimated cost of repairing our beloved Science Center was 200 million dollars. The firm that made the estimate was told to multiply it by 1.85 to make it bigger. We were then told it would be better to build a new science center. Now we learn the new center will be smaller and will cost a billon dollars before the cost overruns. The only win is that the extensive parking lots of the old Science Center will be available for his developer friends.






way way way more people would go into science without the manual lab work.

