@_JasonDerks is presenting at iSCMS! Come by this Saturday afternoon around 2pm to hear his talk: "Increasing proteomics throughput by multiplexing in the mass and time domains". singlecellms.org/?page_id=442
Jason Derks
221 posts

@_JasonDerks
Increasing throughput of proteomics @ParallelSqTech

@_JasonDerks is presenting at iSCMS! Come by this Saturday afternoon around 2pm to hear his talk: "Increasing proteomics throughput by multiplexing in the mass and time domains". singlecellms.org/?page_id=442


The big one is finally out!! In this paper, we set out to provide insight into the fundamental question; How do the individual cells from complex tissues regulate their proteomes? Brief summary of our findings 👇 biorxiv.org/content/10.110…
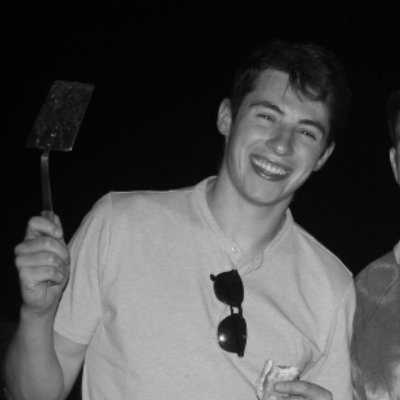

The big one is finally out!! In this paper, we set out to provide insight into the fundamental question; How do the individual cells from complex tissues regulate their proteomes? Brief summary of our findings 👇 biorxiv.org/content/10.110…




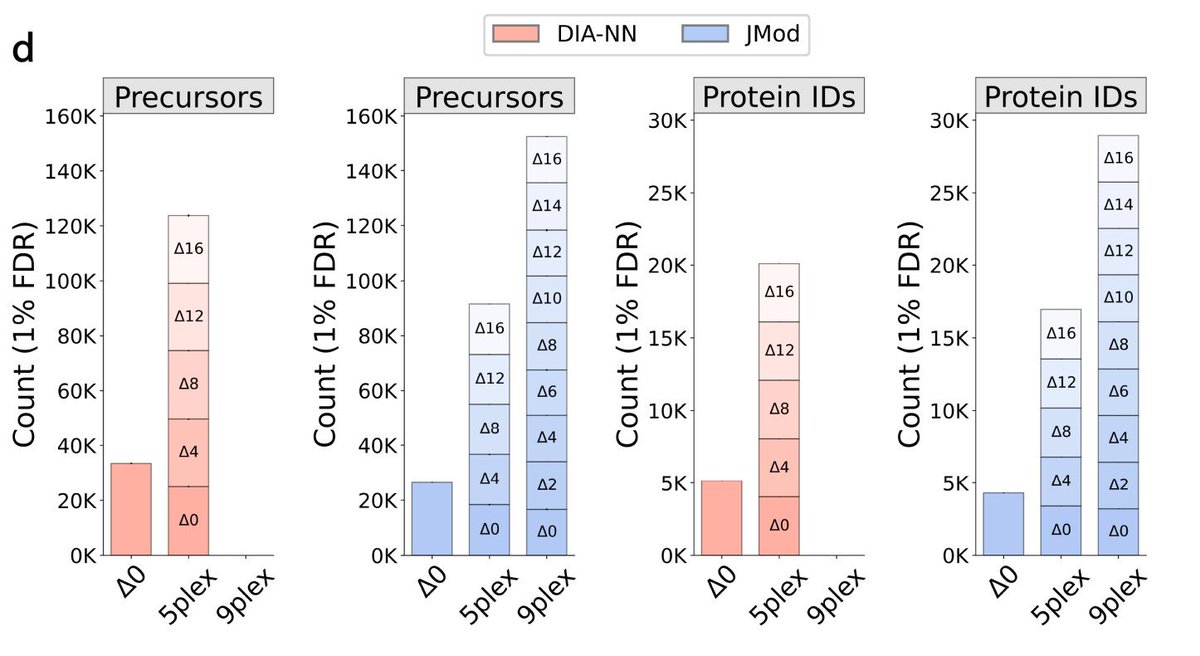
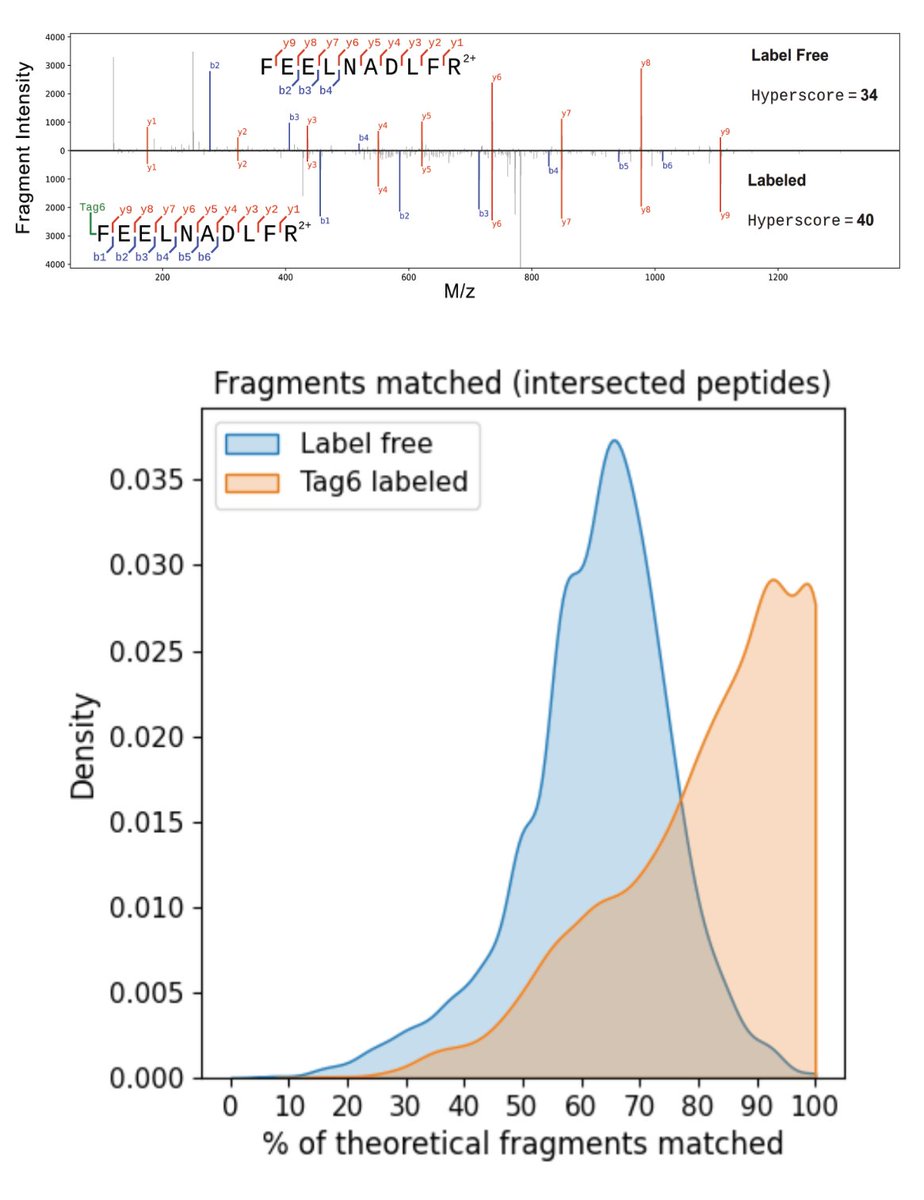
Increasing throughput in mass spec proteomics requires new ways to model the increased complexity of the spectra. To model highly multiplexed mass spectra, we developed JMod. JMod enables new multiplexing approaches in the time and mass domains. biorxiv.org/content/10.110…












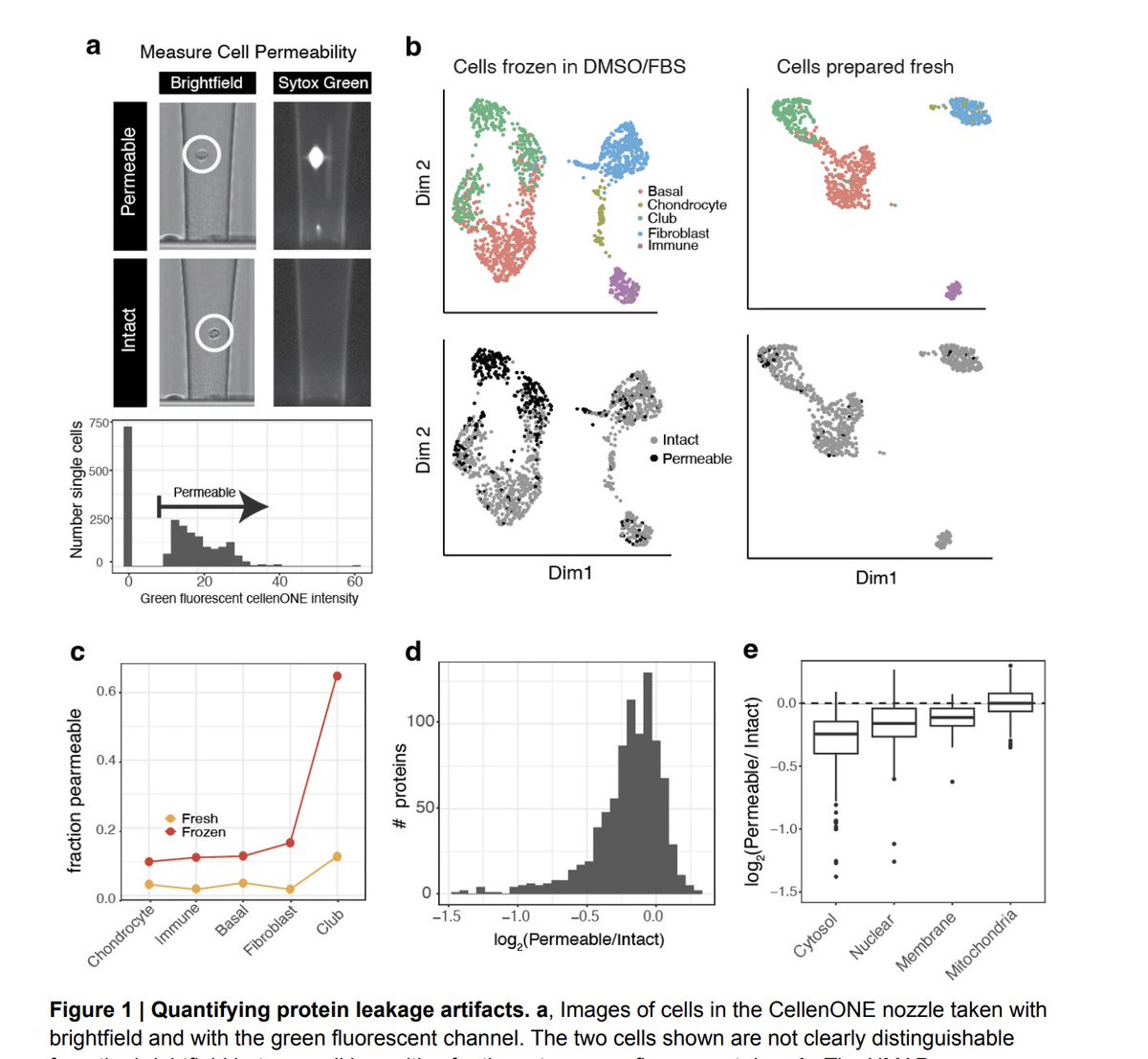
A new preprint: While analyzing proteomes of cells from mammalian tissues, @_AndrewLeduc identified protein leakage from permeabilized cells. We identified clear patterns that can be used to detect & mitigate this issue: biorxiv.org/content/10.110…