Sabitlenmiş Tweet

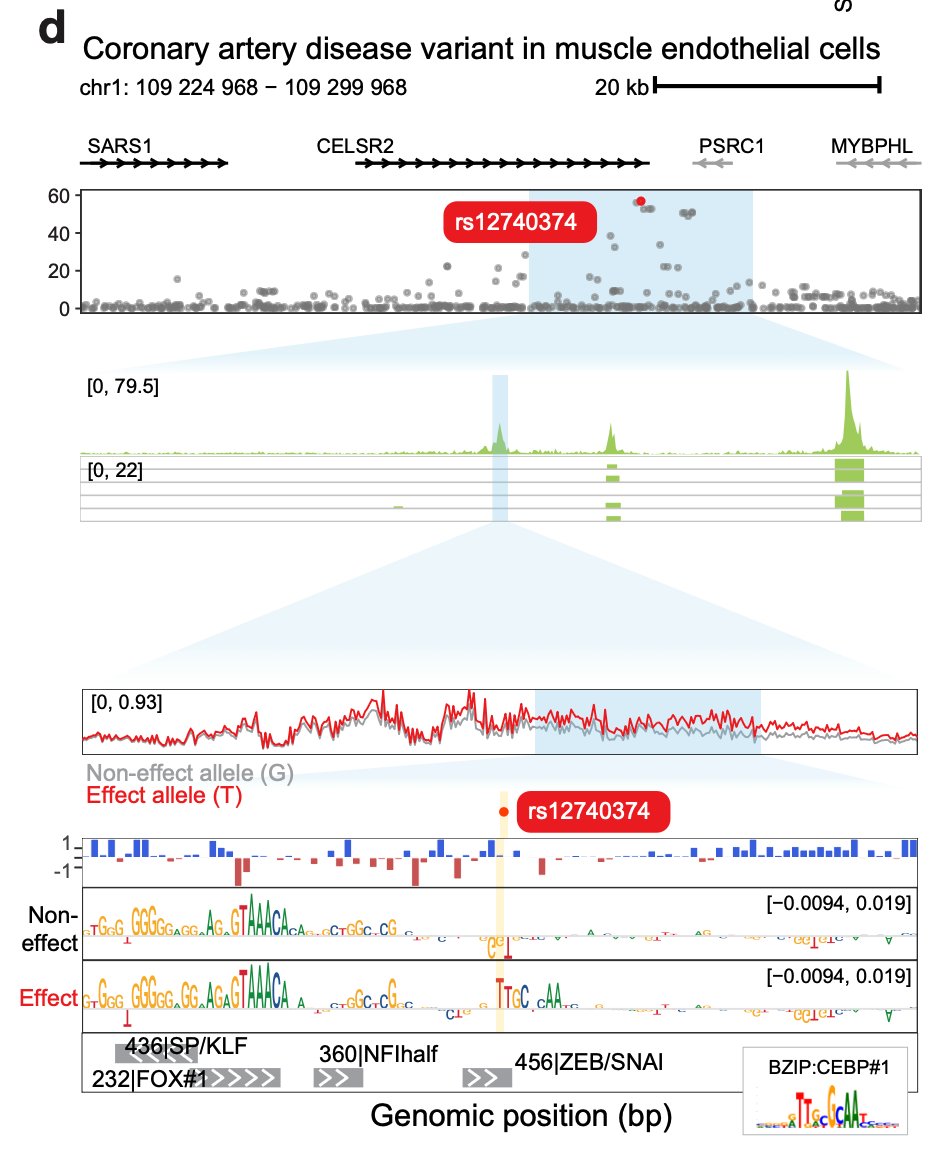
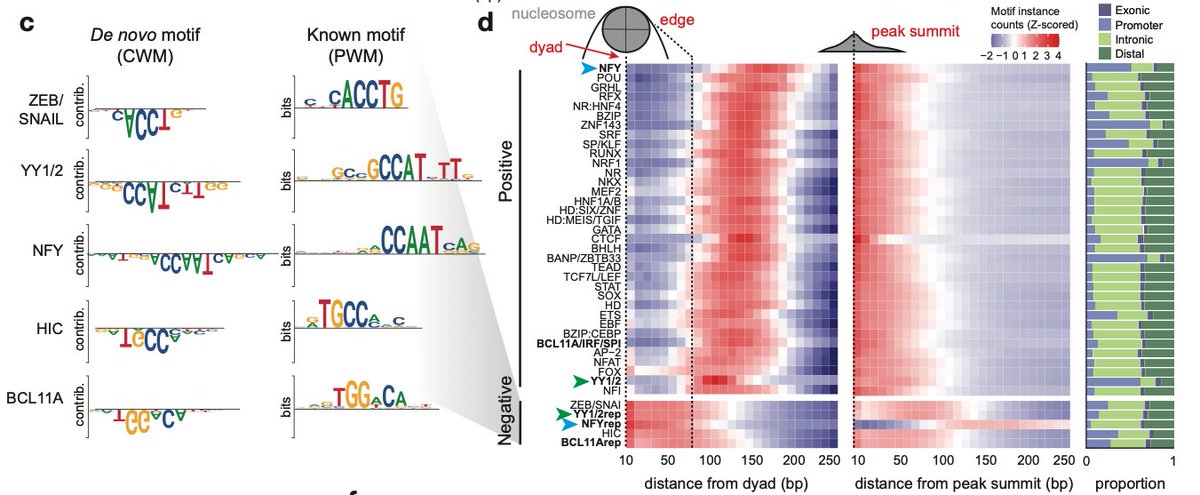
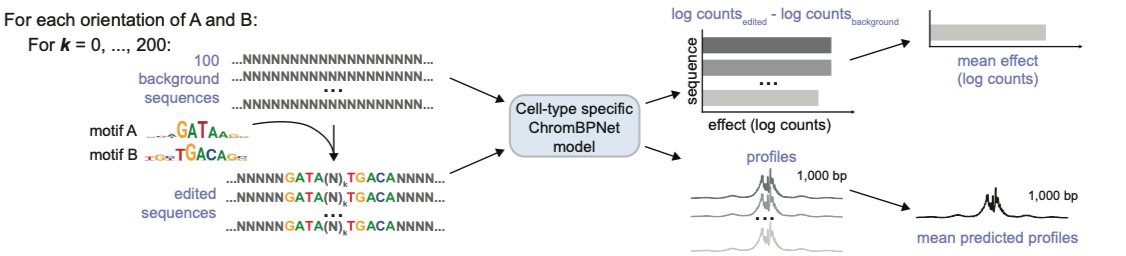
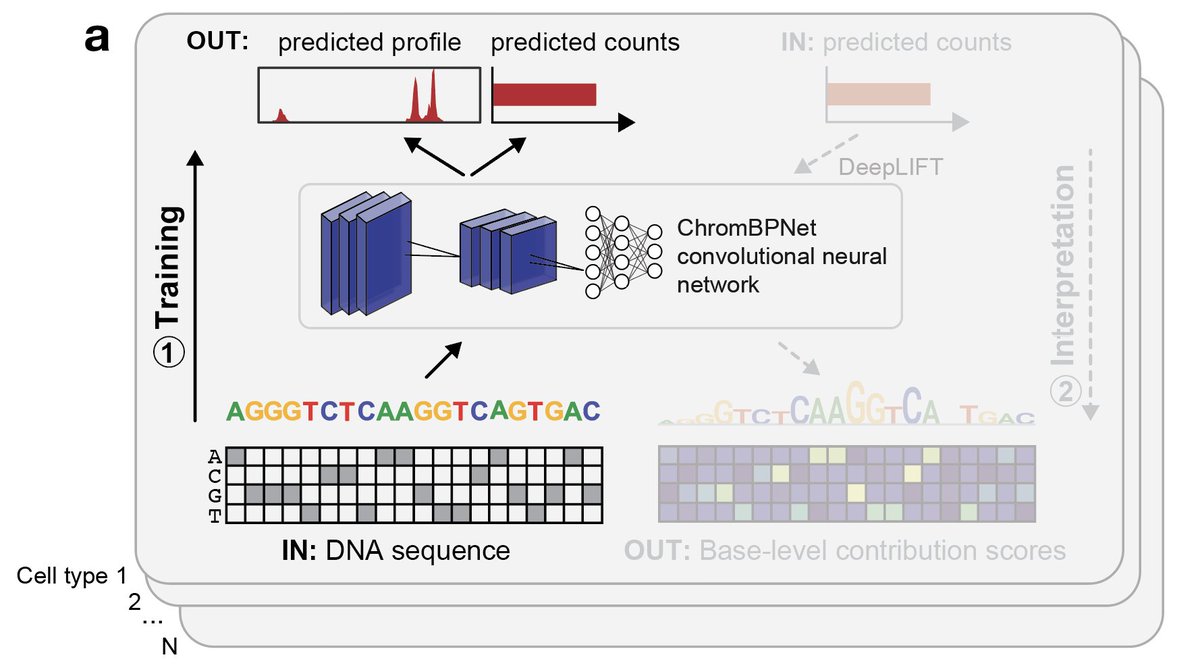
Delighted to share our latest work deciphering the landscape of chromatin accessibility and modeling the DNA sequence syntax rules underlying gene regulation during human development! biorxiv.org/content/10.110…. Read on for more 🧵 [1/16]
English