Benjamin Perry
226 posts

Benjamin Perry
@bots_and_bits
Designing enzymes with bots and bits! | Romero Lab at Duke


This is more evidence that current frontier models remain completely reliant on content-level memorization, as opposed to higher-level generalizable knowledge (such as metalearning knowledge, problem-solving strategies...)

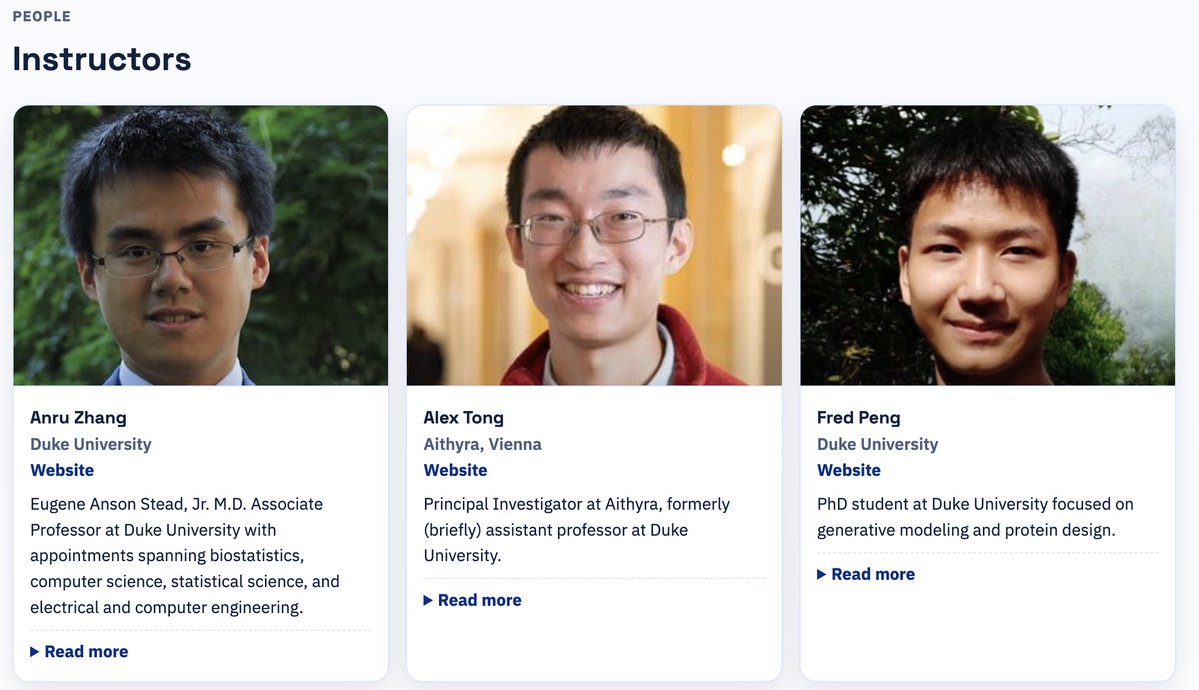
We recently taught a short course at the ENAR 2026 Spring Meeting on generative models for protein, cell, and biomedical data. We’re excited to share the course materials here for anyone interested: pengzhangzhi.github.io/ENAR26-Course-… with @Anru_Zhang, @AlexanderTong7





Using claude code to directly control a liquid handling robot is such a crazy experience


Our paper, “Branched Schrödinger Bridge Matching” (BranchSBM), has been accepted as a main conference paper at #ICLR2026 in Rio! 🌳🧫🇧🇷 In the camera-ready version, we include a new experiment scaling BranchSBM to 11 branches on cell differentiation data! 📷 Check out our freshly updated project page and Github repo below 👇🏻 🌳 Project Page: sophtang.github.io/branch-sbm 📄 Camera-Ready Paper: arxiv.org/abs/2506.09007 💻 Github: github.com/sophtang/Branc… 📹 Reading Group Presentation: youtu.be/inVYA0pQ4Wg?si… Branching is ubiquitous in many dynamical systems, including cell differentiation into distinct fates, diverging cellular responses to drug perturbations, and population dynamics. 🧫 But, existing flow matching and SBM frameworks approximate multi-modal distributions by simulating many independent particle trajectories, which are susceptible to mode collapse, with particles concentrating on dominant high-density modes or traversing only low-energy intermediate paths. To address this challenge, we introduce 🌳 BranchSBM 🌳, a framework that learns a set of diverging velocity fields to reconstruct multi-modal target distributions while simultaneously learning growth networks that allocate mass across branches. 🌳 Our key idea was to define the Branched Schrödinger Bridge Problem as the sum of unbalanced generalized Schrödinger bridge problems, where the weight determines the redistribution of mass across each branch over time. 🌳 We introduce a multi-stage training algorithm to learn the optimal branching drift and growth fields that transport mass along a branched trajectory. This allows BranchSBM to capture diverging, energy-minimizing dynamics without requiring intermediate-time supervision and can generate the full branched evolution from a single initial sample. 🌳 We demonstrate the unique capability of BranchSBM to model dynamic branching trajectories in real-world settings, from differentiating single-cell population dynamics (up to 11 branches!) to simulating diverging cellular responses to drug perturbation. On an unrelated note, I wanted to take this post to congratulate my inspiring and endlessly supportive research mentor, @yinuo_z98, who just defended her PhD and is officially a PhD graduate!! 👩🏻🎓 We’re super excited to present BranchSBM in Rio this April 🇧🇷, along with new workshop papers to be announced! And of course, very grateful for the support from @AlexanderTong7 and @pranamanam 💫

Aaaand it’s online ahhhhh!!! 🥳🥳 So excited!! The first glimpse of my postdoc work with @chorye @dukecagt. Here, @stefanmgolas and I developed TurboPRANCE, an open-source robotics platform for rapid and scaled phage-assisted continuous evolutions. 🧪Tweetorial party!👇1/n



We made FLIP2, a protein fitness benchmark spanning seven new datasets, including enzymes, protein-protein interactions, and light-sensitive proteins, as well as splits that measure generalization relevant to real-world protein engineering campaigns.

Want to fine-tune protein language models but don't have ML experience? 💻❌ We've got you covered! 📢 ✅ Previously: ColabSaprot & ColabSeprot (ESM1/2, ProTrek, ProtBert) 🆕 Now Available: ColabESMC & ColabESM3 Links➡️github.com/westlake-repl/… Tutorial➡️youtube.com/watch?v=nmLtjl…


ColabSaprot & SaprotHub are now in @NatureBiotech! 🧬 A user-friendly, no-code platform for training, sharing, and collaborating on protein language models. We also provide ColabSeprot, integrating ESM1b, ESM2, ProTrek, and ProtBert for the community. nature.com/articles/s4158…


Delighted to share new @arcinstitute work from our group on AI-accelerated lab-in-the-loop, in @ScienceMagazine today One of the most remarkable things about biology is that it's digital. DNA, RNA, proteins: these are all sequences, and their function is directly encoded in their sequence of letters. But a protein of length N has 20^N possible variants and the vast majority are non-functional. Evolution spent billions of years finding the functional needles in this haystack through random exploration and natural selection. For modern biomedicine, we need to solve this in days to weeks.




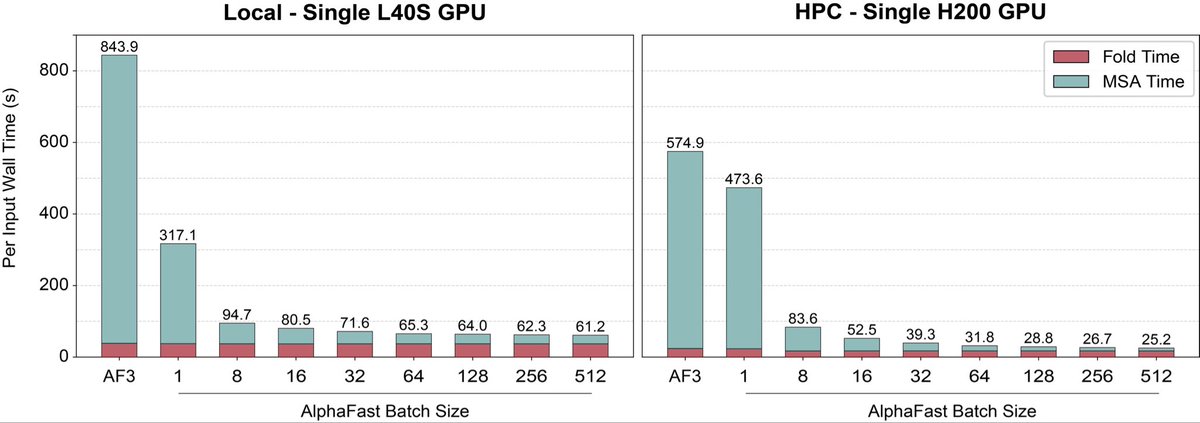
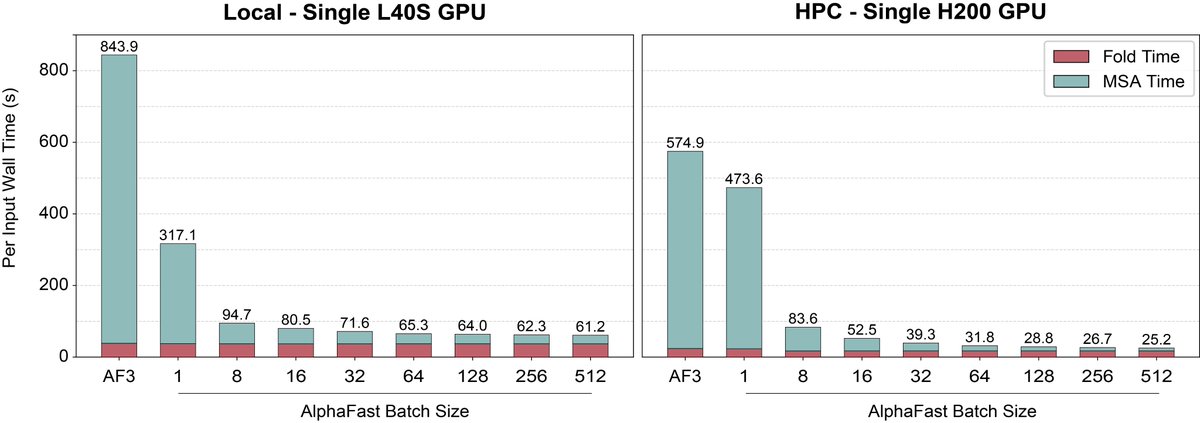
AlphaFold 3 just got a massive speed boost. 🚀 We’re introducing AlphaFast: a GPU-accelerated framework that cuts AF3 inference from >10 mins to ~25 seconds on a single GPU–a 22.8x speedup–without losing structural accuracy. More details below! 1/6 🧵