

Katie Galloway
5K posts


@GallowayLabMIT
Assistant Prof @MITChemE ; mom^4 + wife; enjoys building cell-fate circuits, exploring dna topology, reprogramming the living world, and soccer; soli gloria deo
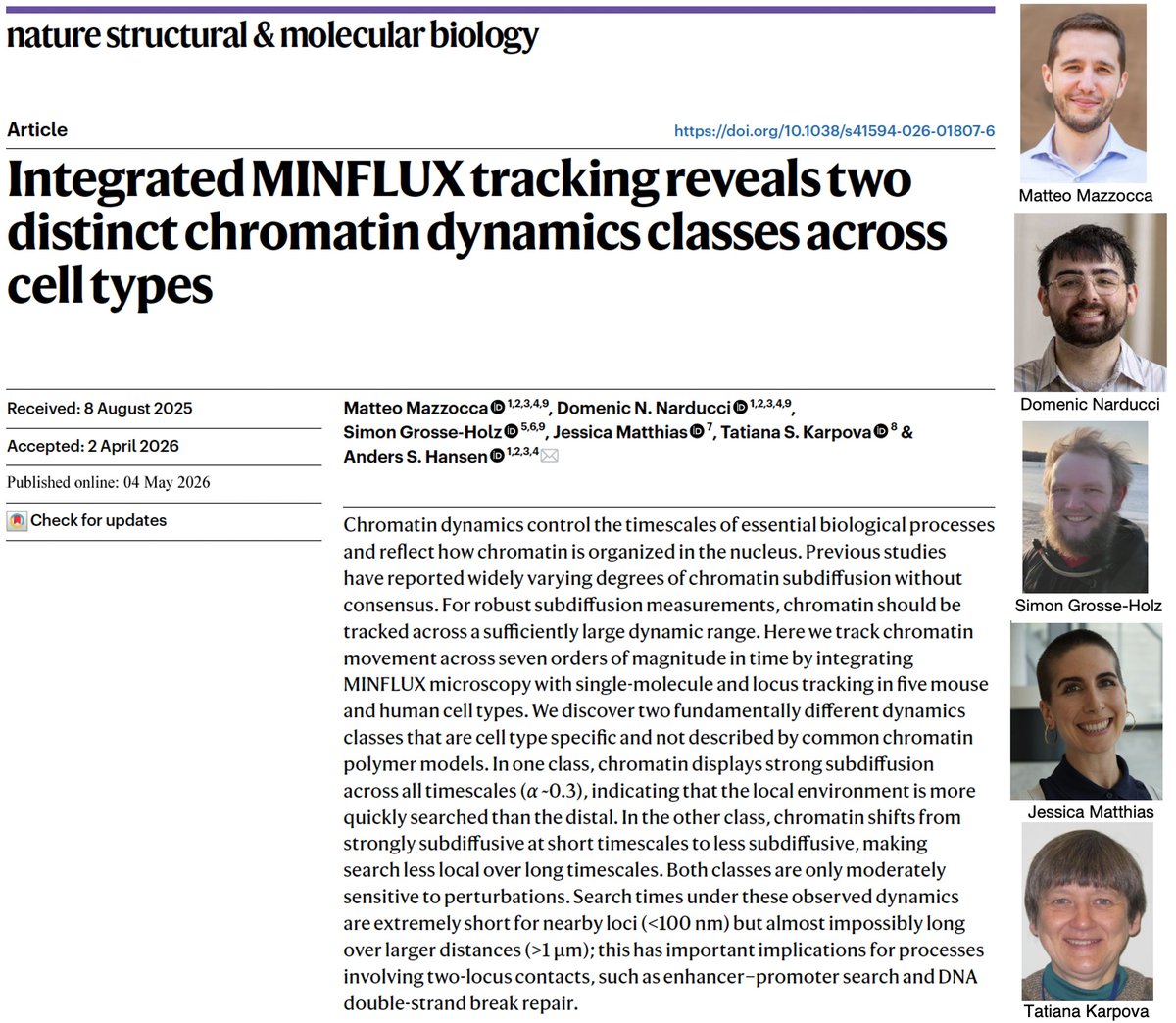






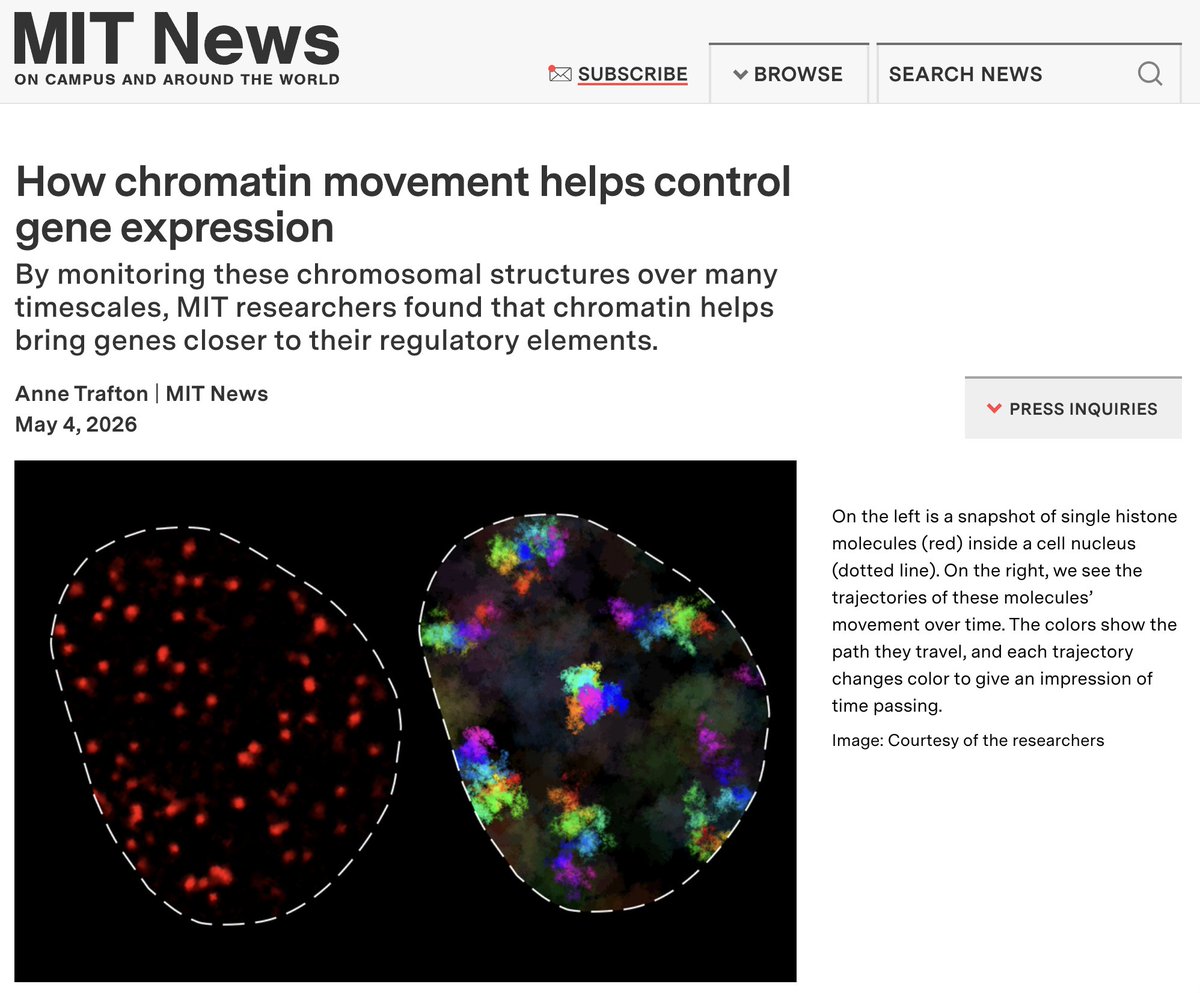
(1/13) Thread on @mazzocca_matteo @DomenicNarducci @SGrosseHolz @_jessematthias new preprint Q: how does chromatin move? Using MINFLUX, SPT & SRLCI, we track chromatin dynamics across 7 orders of magnitude in time to provide some answers biorxiv.org/content/10.110…




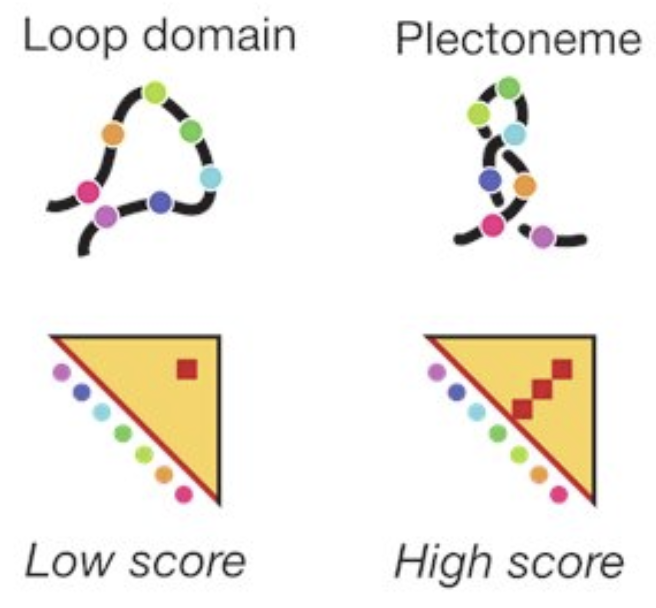
For contact probability, we used Region Capture Micro-C (RCMC) a methods developed by @Anders_S_Hansen's lab which supports targeted pulldown of a region of interest (figure from Goel, V et al. Nature Genetics. 2023) . Since we are just looking at one locus (CLYBL), we want to spend our sequencing dollars getting high resolution at this locus! And boy did we! ( 23/n)
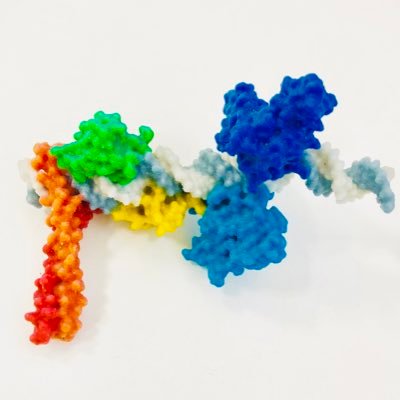

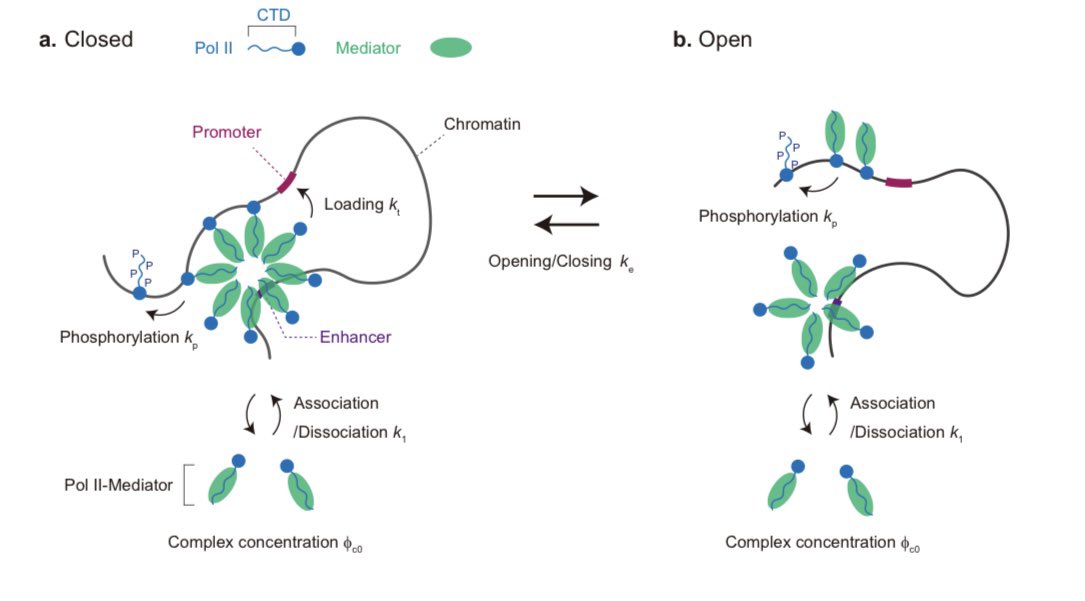









Could the folding of synthetic gene circuits in 3D shape how genes are expressed? Today @ScienceMagazine we report on the role of gene syntax in shaping feedback between transcriptional activity and genome folding for advanced circuit design🧵 (1/n)

STRAIGHT-IN Dual: a platform for dual single-copy integrations of DNA payloads and gene circuits into human induced pluripotent stem cells go.nature.com/4ue2ix4

So you want to engineer your hiPSCs, but targeting DNA payloads requires multiple slow, inefficient steps for each construct. What if we could accomplish multi-site integration seamlessly? Come hear about STRAIGHT-IN Dual now out at Nature Biomedical Engineering! 🧵 Link at end!

Could the folding of synthetic gene circuits in 3D shape how genes are expressed? Today @ScienceMagazine we report on the role of gene syntax in shaping feedback between transcriptional activity and genome folding for advanced circuit design🧵 (1/n)





Stop the doom scrolling! A new 🗞️ from my lab, describing one of our flagship projects of many years we are super excited to share: "Inducible formation of fusion transcripts upregulates haploinsufficient CHD2 gene expression". A 🧵biorxiv.org/content/early/…






Could the folding of synthetic gene circuits in 3D shape how genes are expressed? Today @ScienceMagazine we report on the role of gene syntax in shaping feedback between transcriptional activity and genome folding for advanced circuit design🧵 (1/n)