
ElowitzLab
435 posts

ElowitzLab
@ElowitzLab
Lab of Michael Elowitz at Caltech. Synthetic biology and systems biology





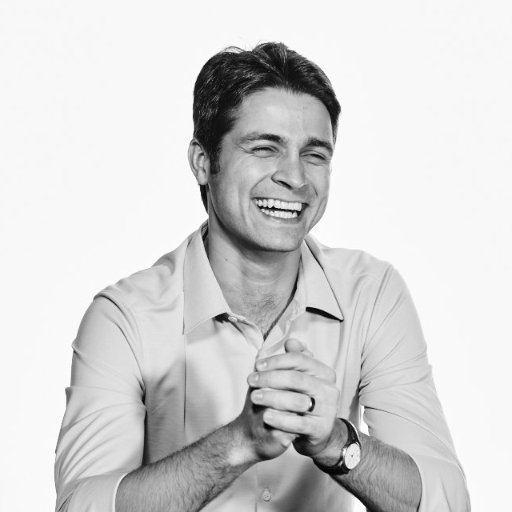
Engineered cells as programmable mRNA delivery vehicles dlvr.it/TSBzt7





Together with Emma Dann, we are thrilled to present a massive new Perturb-seq atlas of 22M primary CD4+ T cells, from 4 donors, across 3 timepoints – the result of a decade-long collaboration between the Marson (@MarsonLab) and Pritchard (@jkpritch) labs. 🧵👇


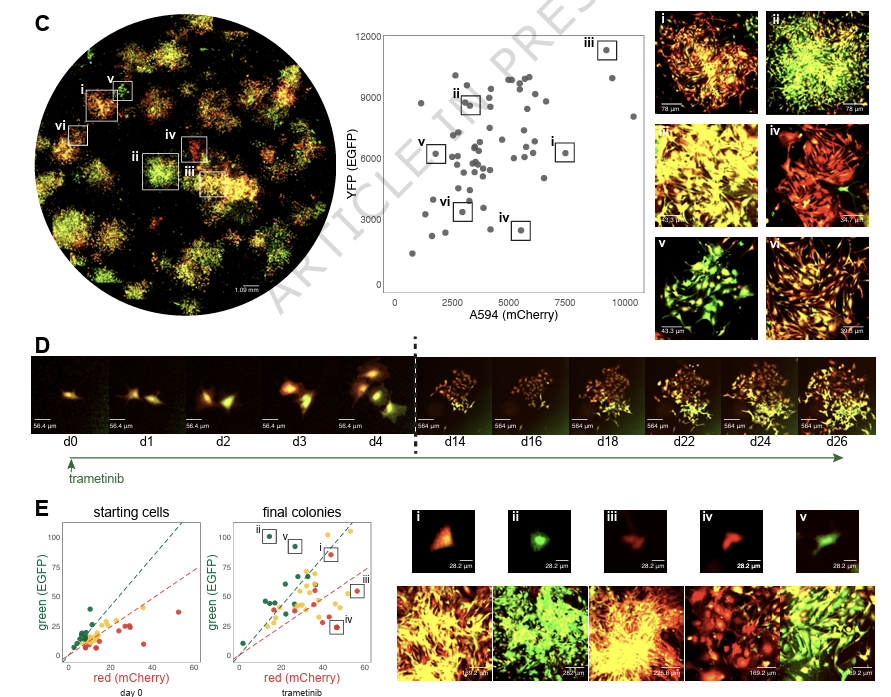
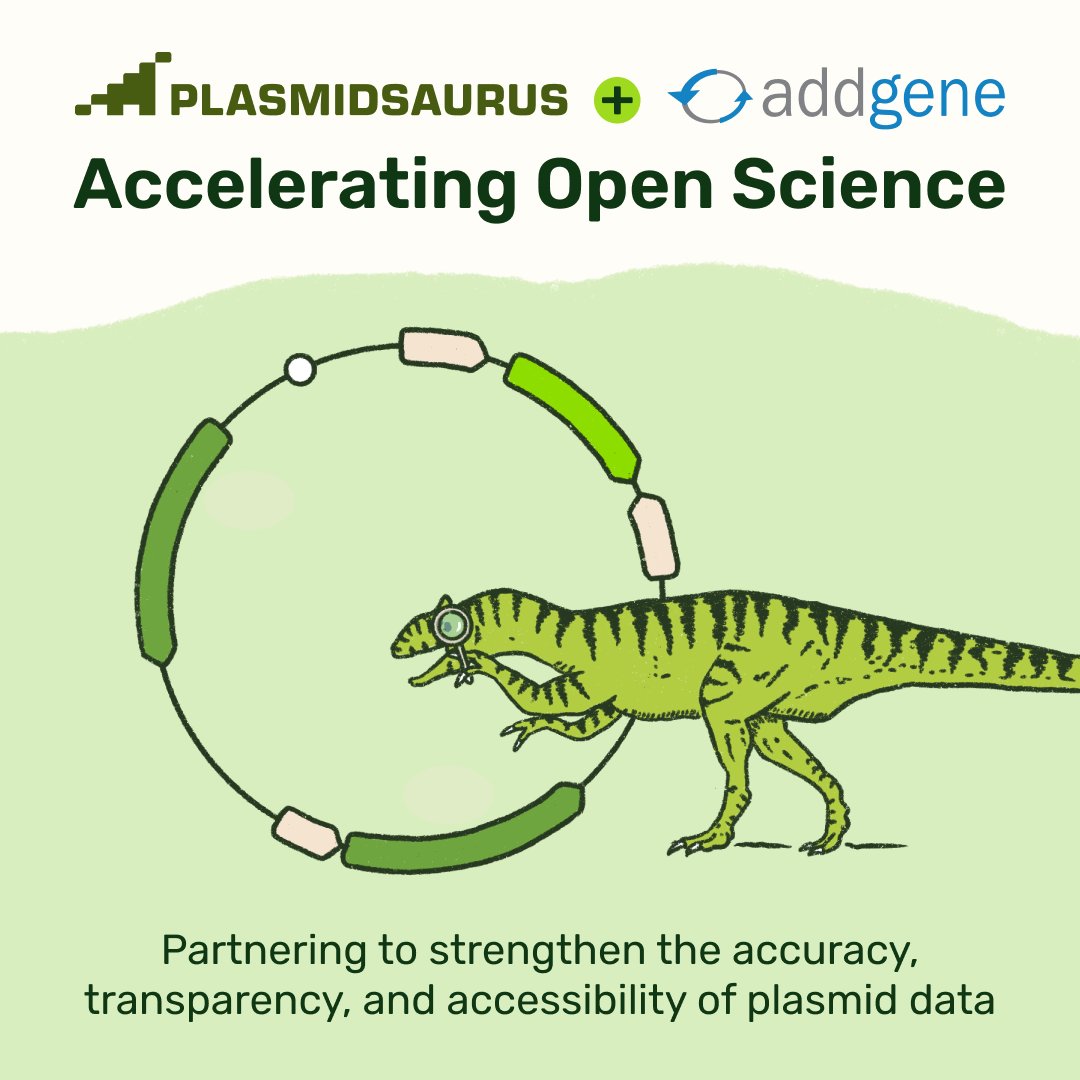
Take two cells and place them side by side. Both cells have the same genome. And yet, oddly enough, they behave in different ways. They divide at different times and their RNA levels are distinct. Now let’s go one step further. Take those same two cells. But this time, imagine that they have not only the same genome, but completely identical molecules at identical concentrations. Will these two cells behave in the same way? The answer is no. This is because there are two types of "noise" inside of living cells; intrinsic and extrinsic. In the first example, the two cells act differently because of subtle differences in their gene levels. Not all genes are expressed at the same time or in the same amount, and this leads to slight differences. This is extrinsic noise, because it is “global to a single cell” but varies “from one cell to another.” In the second example, which is so statistically unlikely as to be basically impossible, the two cells would still have different gene expression patterns “because of the random microscopic events that govern which reactions occur and in what order.” This is intrinsic noise or stochasticity; it is an inalienable part of biology. I’m pulling these quotes from one of my all-time favorite papers, called “Stochastic Gene Expression in a Single Cell.” The first author is @ElowitzLab (of synthetic biology fame) and it was published in August 2022. It’s worth reading. For this paper, Elowitz & co. designed a simple experiment to separate intrinsic and extrinsic noise in a cell. Their goal was measure each source of noise to figure out which one dominates in different scenarios, like exposure to IPTG or the addition of a plasmid. So here’s what they did: First, they took E. coli cells and inserted two genes into the genome; one encoding a fluorescent cyan protein, and another encoding a fluorescent yellow protein. Each gene had the same promoter, and was placed equidistant from the genome’s origin of replication (but on opposite sides.) Next, they grew these cells in LB broth and photographed them using a microscope with color filters. The brightness of each color, in each cell, was quantified. If the variability between different E. coli cells stems from shared cellular conditions (like ribosome levels or extrinsic noise), then both colors in a single cell would fluctuate together. If the variability instead arises from random molecular events (intrinsic noise), then even within the same cell, the cyan and yellow levels would differ. If you plot these changes out on a scatterplot, then you can literally decode which “signals” or “triggers” are dominated by intrinsic or extrinsic noise, and by how much. This is a “beautiful experiment” because the experiment is so simple, yet it retrieves a huge amount of information. All they did was put two genes into an E. coli cell at symmetrical locations in the genome! And from that alone, they deconvoluted noise and its origins.

Our #SynBioFH23 team have brought together an exciting programme to showcase how #bioscience technologies are impacting global healthcare approaches. #GlobalHealth Submit an abstract by 17 Jan 2023 for an opportunity to share your work.📅 Full details➡️bit.ly/3Dj2HHp


I'm excited to share our new preprint on LagTag, a method that recovers both past and present chromatin states from the same mammalian cells. biorxiv.org/content/10.110…




