Chris Hayduk
1.9K posts

Chris Hayduk
@ChrisHayduk
Writing About AI, Biology, & Geopolitics on Substack || Machine Learning Engineer @Meta || Prev. Lead Machine Learning Engineer for Drug Discovery @Deloitte






$WMT is disappointed in results from OpenAI partnership, whereby Walmart users are allowed to shop via ChatGPT and OpenAI would receive a commission on these purchases “Conversion rates—the percentage of users following through with a purchase of an item shown to them by ChatGPT—have been three times lower for the selection sold directly inside the chatbot than those that require clicking out, according to Daniel Danker, who oversees design and product for Walmart. Put simply, Instant Checkout has been a flop.” -- Wired OpenAI on a heater recently in the news….

Universal embeddings FTW 😊 One of the flagship projects at FAIR was to "embed the world" (i.e. represent every entity on Facebook). The name was soon changed to "Filament", deployed internally, and eventually open-sourced as "PyTorch-BigGraph" The techniques were more primitive than today! github.com/facebookresear…


I understand and agree with many criticisms of philanthropy. But practically, fortunes have to go somewhere. There are only 3 options: philanthropy, heirs & govt. If not nonprofits, is Peter Thiel's plan to give $10B+/child? I'm more skeptical of that than he is of philanthropy.


I'm delighted to announce that @_MathAcademy_ has released two courses in Mathematical Methods for the Physical Sciences. Designed for students who want the mathematical tools needed for undergraduate-level study in physics, engineering, and other STEM fields. Details below👇


Austin, TX from the same spot 10 years apart (2014 vs 2024) Pretty stunning transformation.


1/ Introducing NanoGPT Slowrun 🐢: an open repo for state-of-the-art data-efficient learning algorithms. It's built for the crazy ideas that speedruns filter out -- expensive optimizers, heavy regularization, SGD replacements like evolutionary search.

















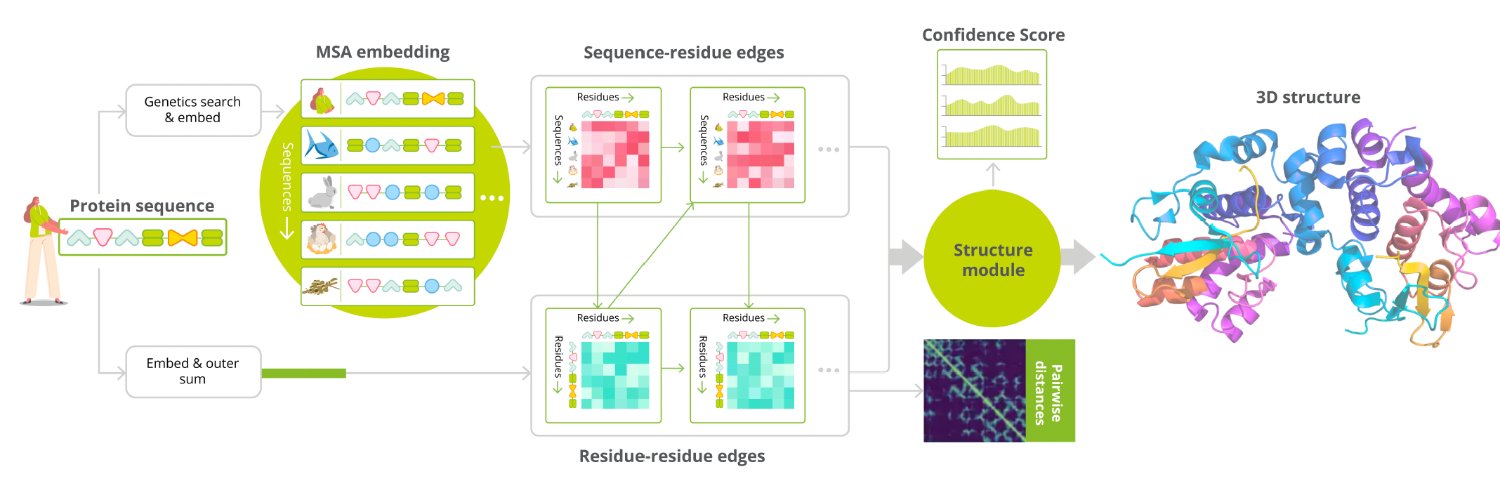
I'm rebuilding AlphaFold2 from scratch in pure PyTorch. No frameworks on top of PyTorch. No copy-paste from DeepMind's repo. Just nn.Linear, einsum, and the 60-page supplementary paper. The project is called minAlphaFold2, inspired by Karpathy's minGPT. The idea is simple: AlphaFold2 is one of the most important neural networks ever built, and there should be a version of it that a single person can sit down and read end-to-end in an afternoon. Where it stands today: - ~3,500 lines across 9 modules - Full forward pass works: input embedding → Evoformer → Structure Module → all-atom 3D coordinates - Every loss function from the paper (FAPE, torsion angles, pLDDT, distogram, structural violations) - Recycling, templates, extra MSA stack, ensemble averaging — all implemented - 50 tests passing - Every module maps 1-to-1 to a numbered algorithm in the AF2 supplement The Structure Module was the most satisfying part to build. Invariant Point Attention is genuinely beautiful — it does attention in 3D space using local reference frames so the whole thing is SE(3)-equivariant, and the math fits in about 150 lines of PyTorch. What's next: - Build the data pipeline (PDB structures + MSA features) - Write the training loop - Train on a small set of proteins and see what happens The repo is public. If you've ever wanted to understand how AlphaFold2 actually works at the level of individual tensor operations, this is meant for you. Repo: github.com/ChrisHayduk/mi…




