GAMA Miguel Angel 🐦⬛🔑
4.4K posts


GAMA Miguel Angel 🐦⬛🔑
@miangoar
Biologist that navigates in the oceans of diversity through the space-time | MSc in Biochem/Bioinfo @ibt_unam 🇲🇽 | Protein evo, metagenomics & AI/ML/DL












“In the long run, interpretability is hopeless. Physics being reducible to equations was an exception, there is no reason for the world to be comprehensible to our puny brains.” – @MoAlQuraishi #ICLR2024

Today we announced our $2.1 Billion funding. This is the catalyst that will bring us to a future of medicine with the power of AI. And we are just getting started, come join our mission.



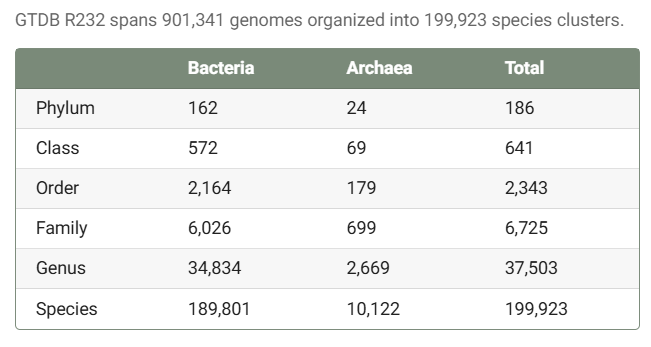
It’s estimated that the Protein Data Bank (PDB) cost around $13B to create. Alphafold was only possible because of it. If we want ML to solve biology, we should be funding the creation of databases and the development of new assay technologies. ML is nothing without data.


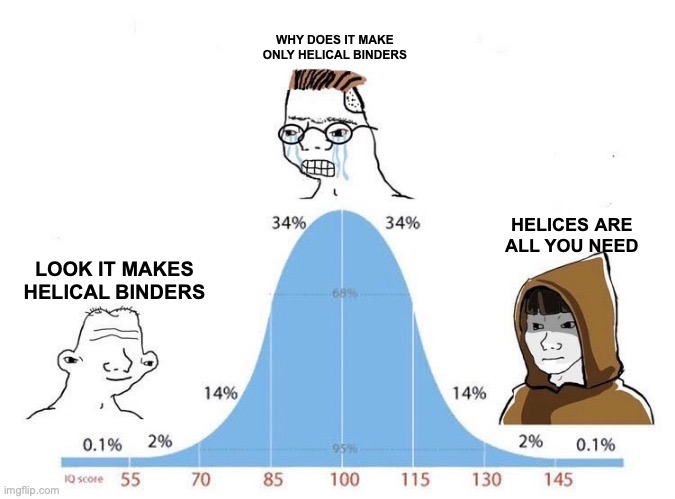
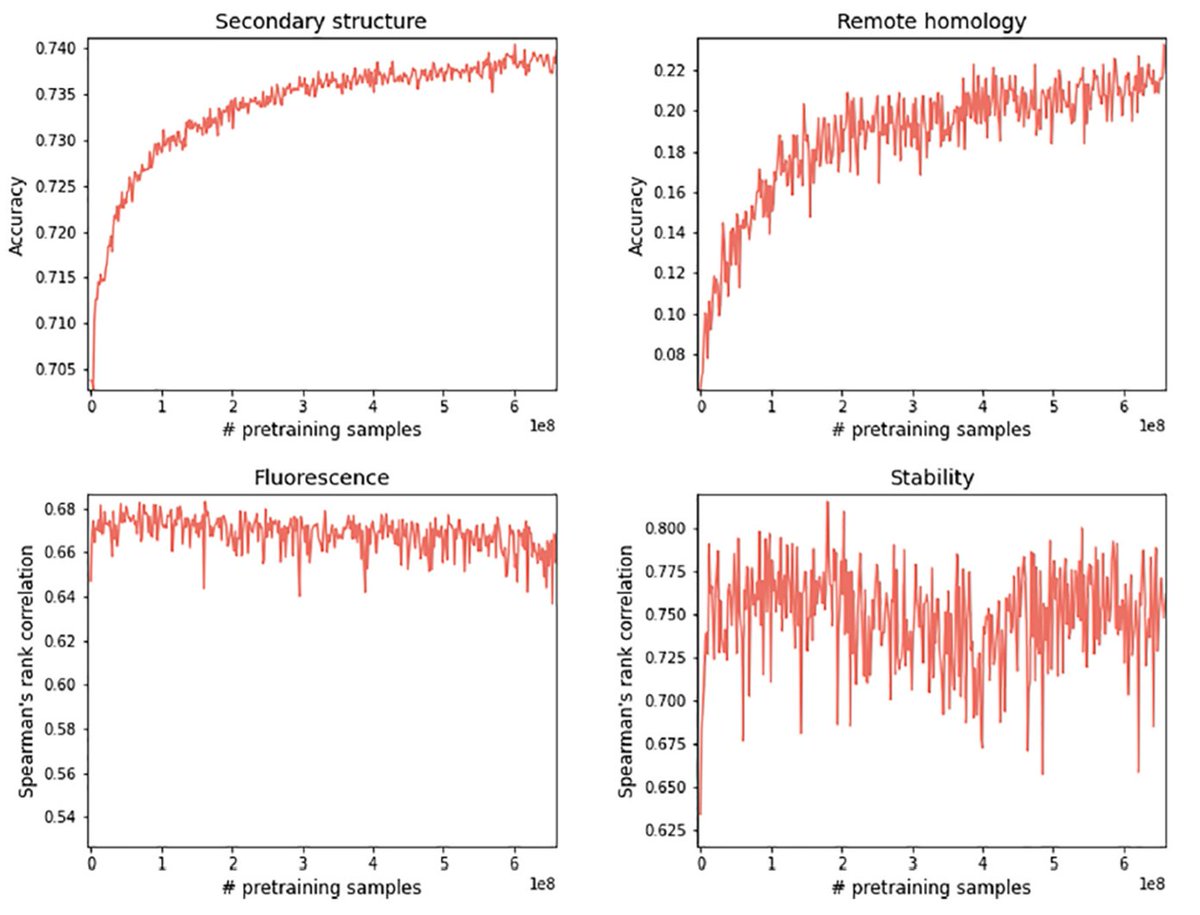
1/2 For zero-shot fitness prediction, protein language models hit a wall around Spearman correlation values of ~0.5. But this isn’t because the models are trash it’s because the protein universe is highly biased and PLMs overfit to the most abundant protein families.







Introducing Genie 3, a generative protein model that substantially advances the state-of-the-art for binder design, increasing in silico success rates by up to 20x on hard multimeric targets. It also debuts a form of inference-time scaling unobserved in other design models. 🧵1/8