IntoWild
1.8K posts






Ever wonder why we decide to build a new microscope and how we go about it? For the last lecture this semester of the graduate microscopy class Na, Gokul, and I teach at Cal, I spoke on our developing iSOAR microscope, a swept HILO design with adaptive optics and structured illumination. iSOAR is designed to produce the petabytes of high resolution 5D videos of subcellular dynamics in zebrafish we need to train the multimodal vision models at the heart of our Cell Observatory Initiative. We still have some work to do, but the scopes are coming along nicely. It's an exciting time at the Observatory, so if you're experienced and looking for a job in zebrafish transgenics, ultra-high throughput image processing, or the development of 5D spatiotemporal foundation models for interpreting the insane dynamic complexity of living matter, we'd love to hear from you. (sup@berkeley.edu, betzige@janelia.hhmi.org).

A new @IMol_Institute study in Science Advances sheds light on mitochondrial protein import. science.org/doi/epdf/10.11… Huge credit to this outstanding team of co-authors!
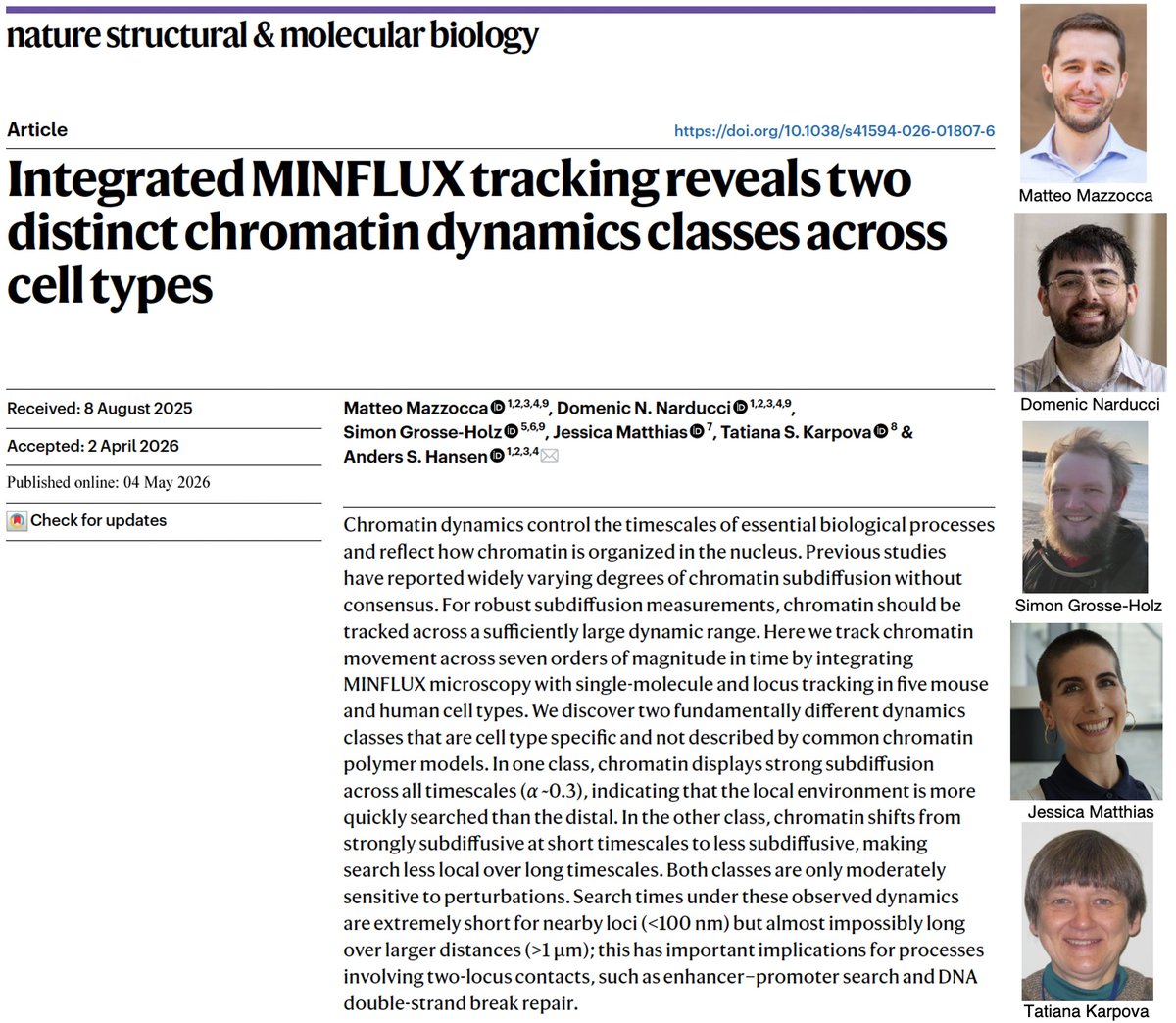


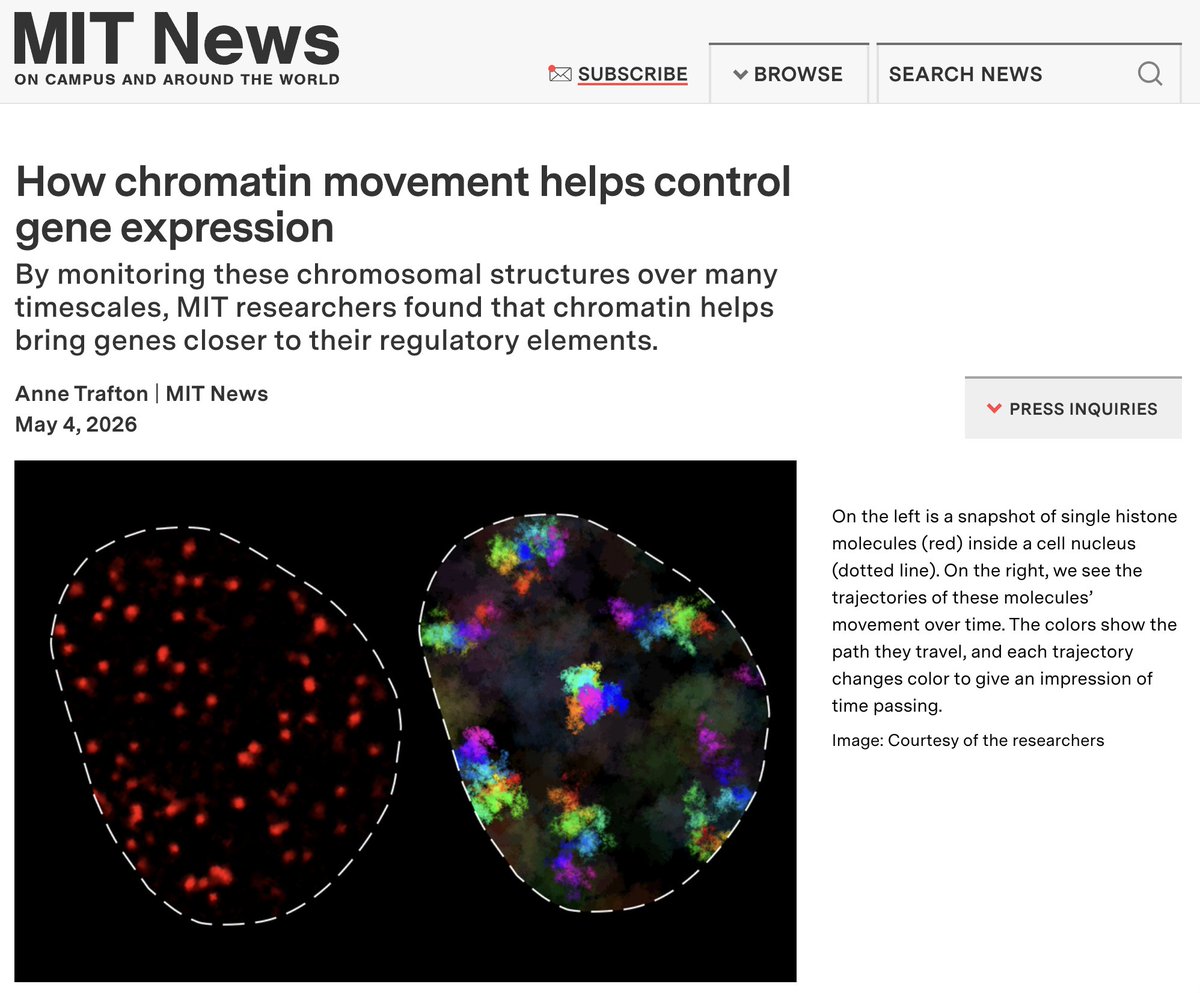
(1/13) Thread on @mazzocca_matteo @DomenicNarducci @SGrosseHolz @_jessematthias new preprint Q: how does chromatin move? Using MINFLUX, SPT & SRLCI, we track chromatin dynamics across 7 orders of magnitude in time to provide some answers biorxiv.org/content/10.110…




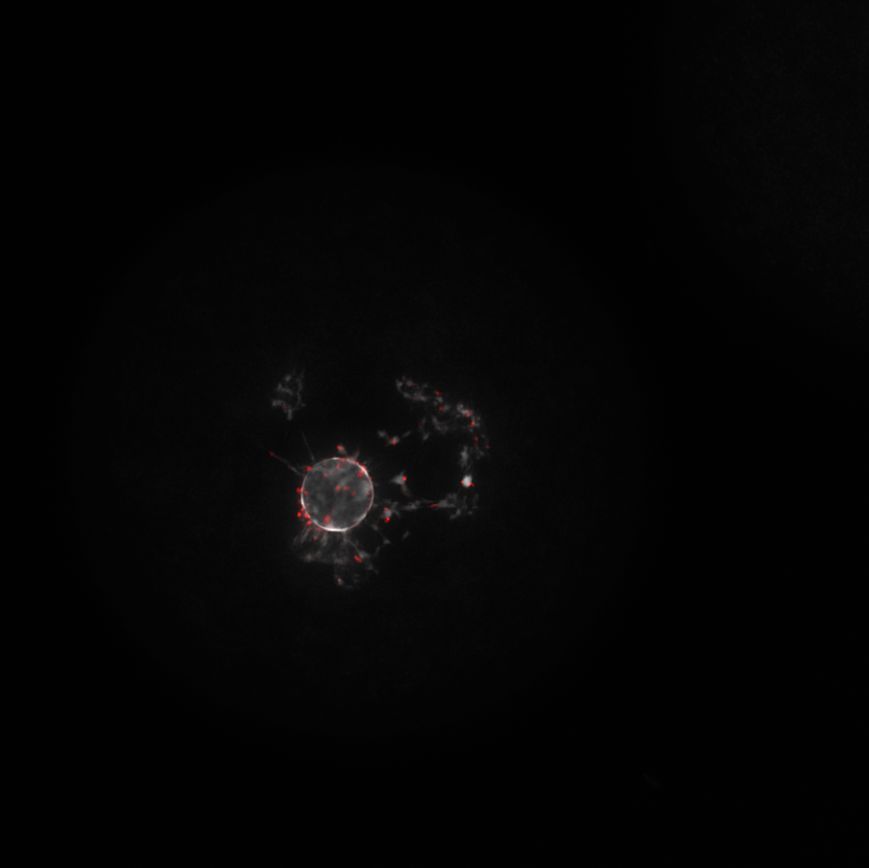


Our live tissue clearing paper is out in @naturemethods! We achieved optical clearing of mammalian brain tissues without compromising normal neuronal function. Big congrats to @Shigenori774 and our wonderful collaborators! 🎉 nature.com/articles/s4159… (1/10)