Sabitlenmiş Tweet

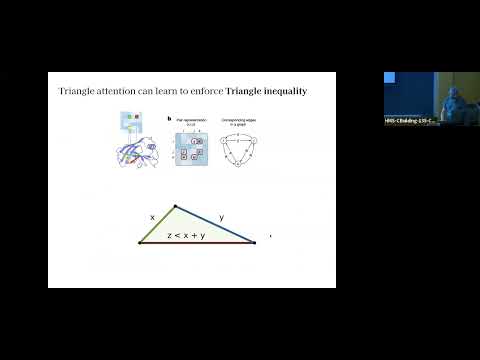
CG models can be quickly (~s) and very accurately (<0.5 A, ~2 MolProbity score) reconstructed to atomistic structures using ML.
"One particle per residue is sufficient to describe all-atom protein structures" biorxiv.org/content/10.110…
English