immuneML
59 posts


immuneML
@immuneml
immuneML is a software platform for machine learning analysis of adaptive immune receptors and repertoires | tweets by @milenapavl and @victorgreiff


Finally biologists can also use numpy (array programming). Handling e.g. DNA and protein sequences with convenience and speed, like physicists and machine learners for decades have worked with numerical data: nature.com/articles/s4159… (1/3)



(1/8)🎉A fresh preprint in which we present LIgO — a powerful tool to simulate adaptive immune receptor (AIR) and repertoire (AIRR) data for the development and benchmarking of AIRR-based ML 🧵⬇️ biorxiv.org/content/10.110…

1/8 New preprint: generative modeling of AIRR repertoires, the last piece of my PhD, >2 years of work, a project that is very dear to me biorxiv.org/content/10.110…

Happy to share my first preprint in the Immunolingo project “Advancing protein language models with linguistics: a roadmap for improved interpretability”, a perspective on adapting LMs for protein sequences with the help of linguistic knowledge. arxiv.org/abs/2207.00982

While R is succinct and intuitive for numbers, I have mostly found it cumbersome for use with text or biosequence data. With the new BioNumPy library (biorxiv.org/content/10.110…), working with sequence data in Python feels like any other (numeric) vectors/matrices/arrays. 1/3

With BioNumPy, I claim my postdocs @ivargrytten and Knut Rand have given the entire bioinformatics community a Christmas present🫢☺️. This is the fundamental library for working effectively with sequence data that I have been missing since I entered bioinformatics 17 years ago!


. @pandaisikit, @probertimmodels, @victorgreiff and colleagues introduce the Absolut! framework, which can generate synthetic 3D-antibody-antigen structures to assist machine learning and dataset construction for antibody design. nature.com/articles/s4358… 👉rdcu.be/c1UcJ


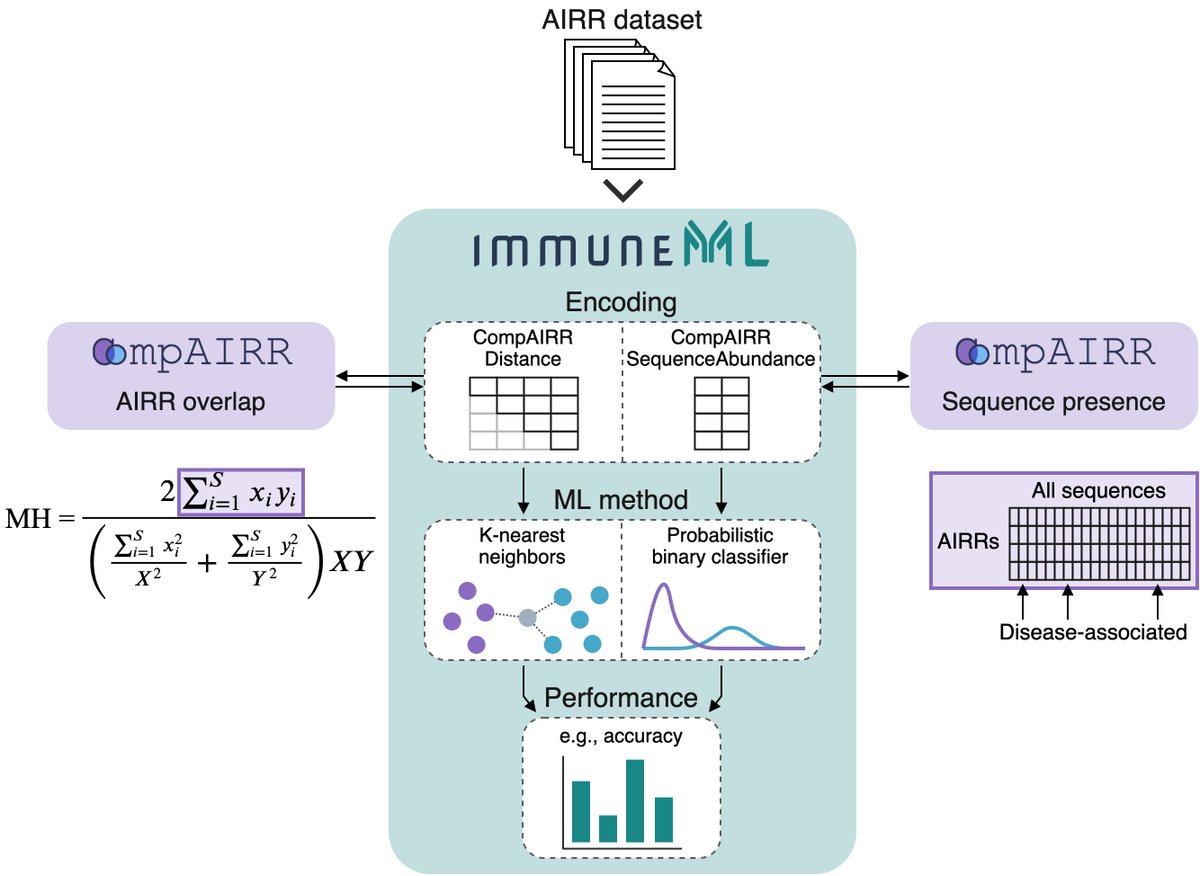
We just released CompAIRR – a tool that reduces greatly the longstanding computational bottleneck of identifying shared identical and similar sequences among large sets of immune repertoires. All the details/links are in the thread below. Feedback is very welcome! @airr_community




Out now in @GigaScience after substantial improvement since the first version: doi.org/10.1093/gigasc…. Improvements since the first version are briefly summarised in the 🧵