Geir Kjetil Sandve
227 posts

Geir Kjetil Sandve
@SandveGeir
Professor in bioinformatics/machine learning at University of Oslo, Norway. Follow me at @[email protected] or at @[email protected]



1/12 🧵 Quick thread: a heads-up for anyone working with high-dimensional omics data and multiple testing — FDR correction can sometimes lie to you. A thread based on our recent publication in @GenomeBiology : doi.org/10.1186/s13059…
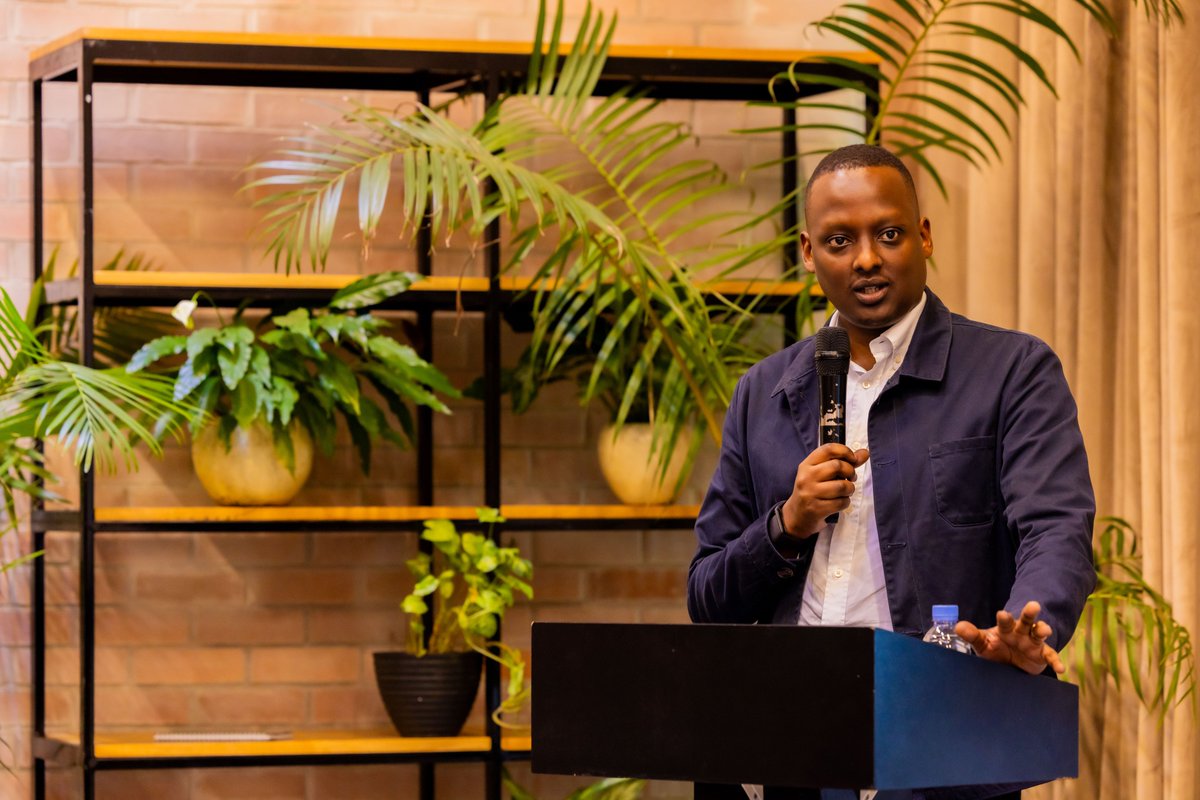
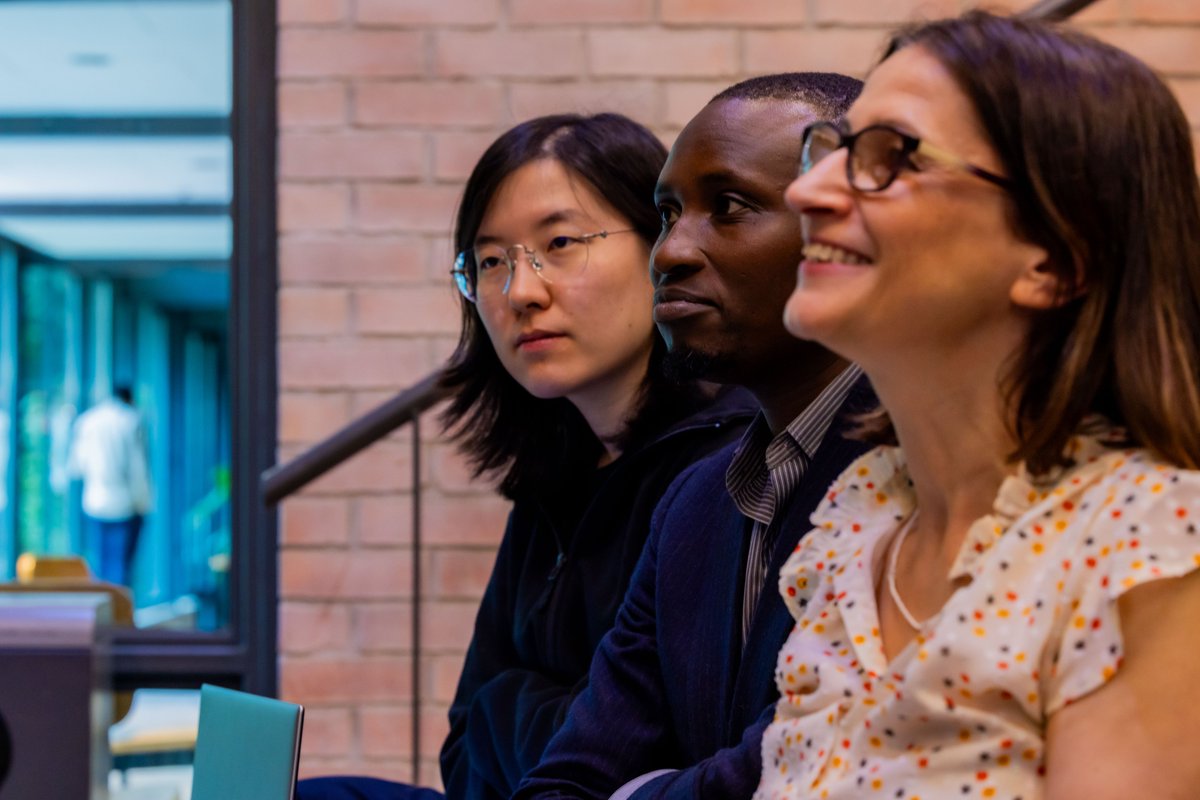
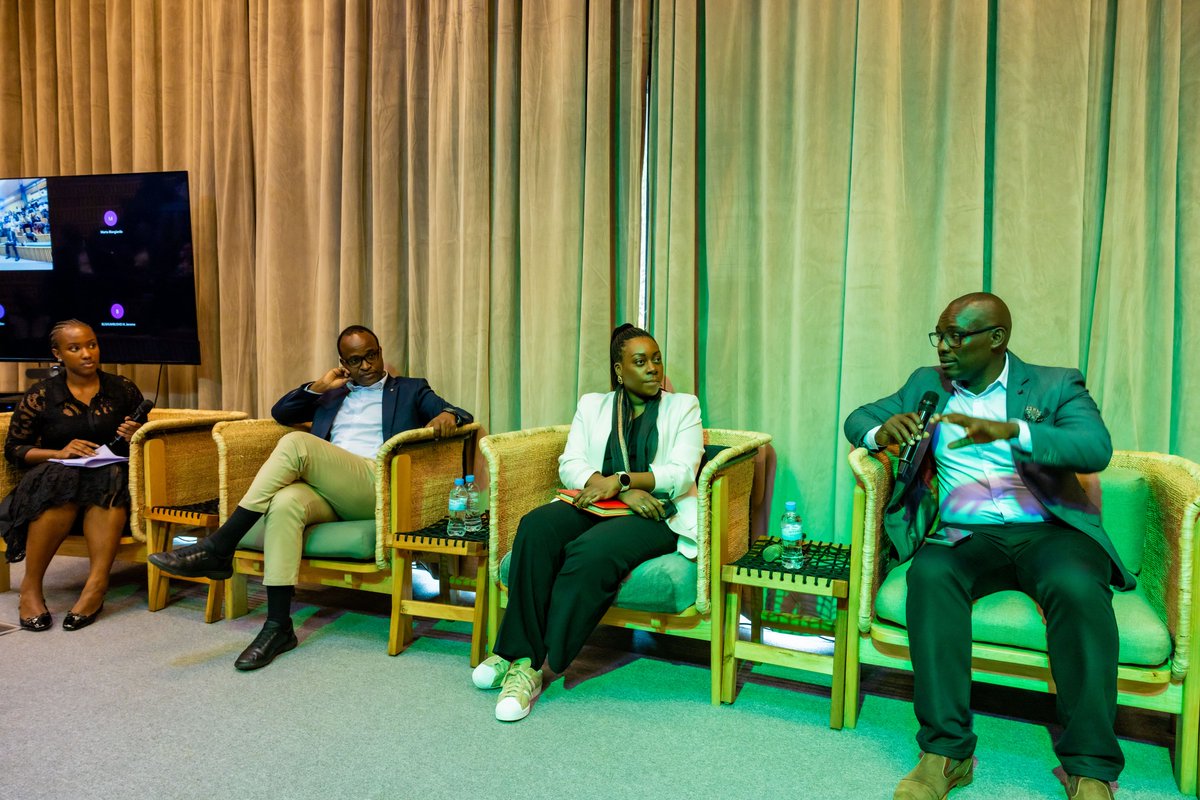















(1/10) Our work on antibody developability is out! We present a cartography of the developability landscapes of the native and human-engineered antibodies. This study was a team effort of @HabibBashour @EvaSmorodina @pariset_matteo , and @Jahn_Zhong biorxiv.org/content/10.110…

📢 @maispaceha, @PRobertImmodels, @BartlomiejSwia4, @SandveGeir, @victorgreiff et al. explore the potential of linguistic formalization of antibody language to improve rule mining in deep LMs for antibody sequence analysis. nature.com/articles/s4358… ➡️rdcu.be/dKOdq

📢 @maispaceha, @PRobertImmodels, @BartlomiejSwia4, @SandveGeir, @victorgreiff et al. explore the potential of linguistic formalization of antibody language to improve rule mining in deep LMs for antibody sequence analysis. nature.com/articles/s4358… ➡️rdcu.be/dKOdq


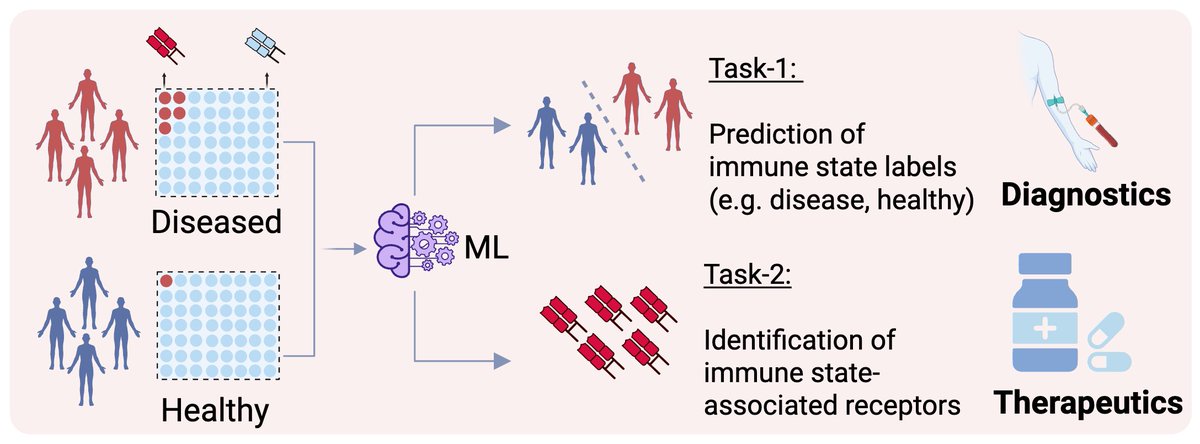
Our perspective on the generalization of machine learning-identified biomarkers is now published in @NatMachIntell 🎉 nature.com/articles/s4225…

KAGE2 is out! Enables very fast and accurate genotyping of structural variants using pangenomes: biorxiv.org/content/10.110…. I’ve spent the last 6+ months going deep into the SV rabbit hole, and had some surprises I thought it’s worth to also share (1/6)






Happy to share my first preprint in the Immunolingo project “Advancing protein language models with linguistics: a roadmap for improved interpretability”, a perspective on adapting LMs for protein sequences with the help of linguistic knowledge. arxiv.org/abs/2207.00982