Eugene Katrukha 🇺🇦
3.9K posts


Eugene Katrukha 🇺🇦
@katpyxa
biophysics, cytoskeleton and self-disorganization
Utrecht, NL Katılım Eylül 2008
721 Takip Edilen1.8K Takipçiler
Eugene Katrukha 🇺🇦 retweetledi

What is the difference between a growing and shortening microtubule end? I thought I knew the answer until I started talking to @maxotubule and @KalutskiyMaksim. Turns out the key are protofilament clusters - check out our preprint for details biorxiv.org/content/10.110…

English
Eugene Katrukha 🇺🇦 retweetledi

Have you ever analyzed a #microscope image before- even just a single one? Please take our survey on progress and places to still improve for image analysis software! #microscopy #imageanalysis #bioimageanalysis forum.image.sc/t/2024-image-a…
English
Eugene Katrukha 🇺🇦 retweetledi

.@yuhao_han, Grünewald, @MarinaMikhaylo8 @HumboldtUni et al. use primary hippocampal cultures to demonstrate that neurons with axon-carrying–dendrite (AcD) morphology can develop independent of an in vivo environment. hubs.la/Q02WP-Fw0
@christlet #Neuroscience #Polarity
English
Eugene Katrukha 🇺🇦 retweetledi
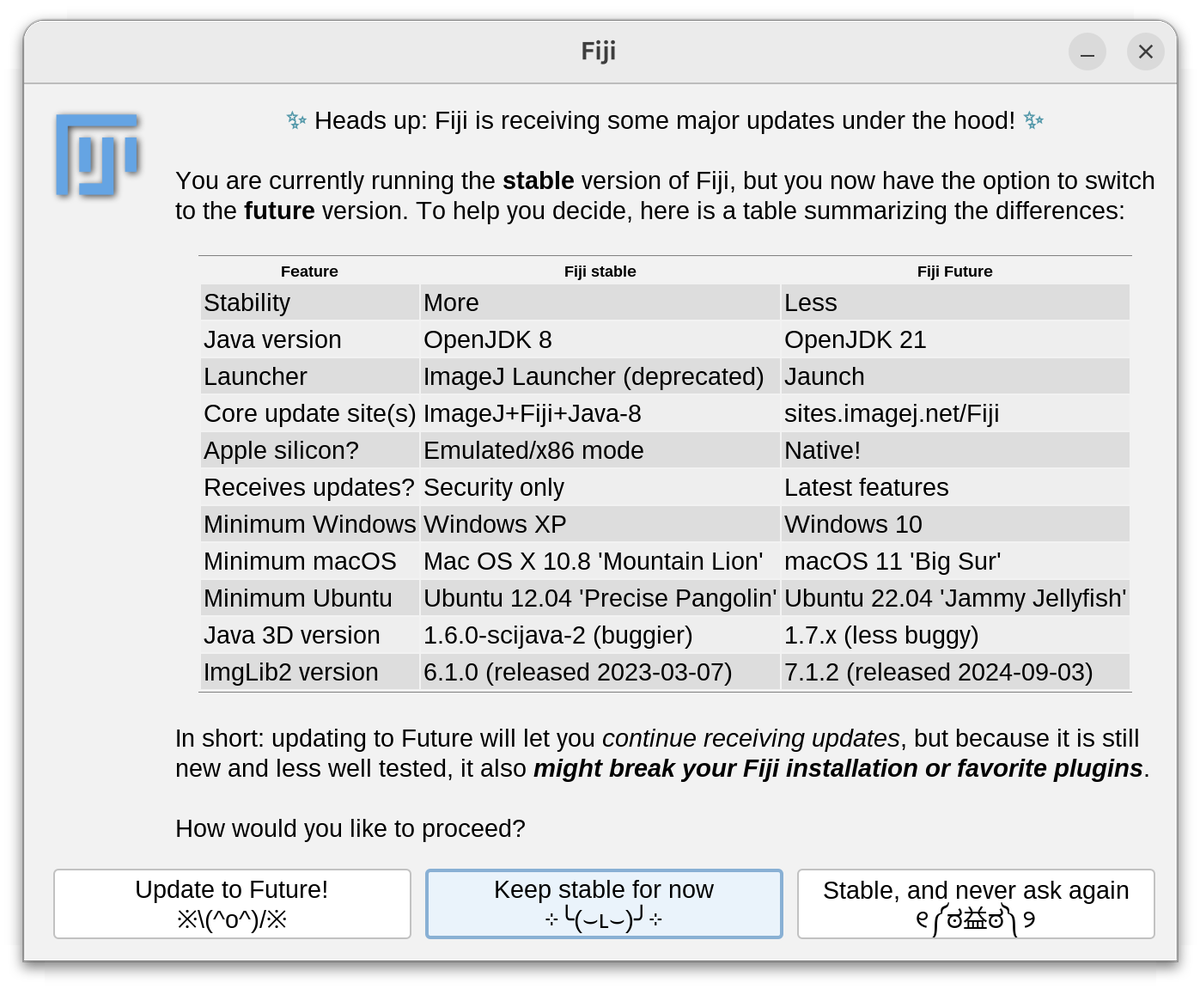
@christlet @ArrateMunoz @guijacquemet Pre-release bundles available at downloads.imagej.net/fiji/future/ – still working on fixing a few things, which is why I haven't pushed the option to update to all Fiji users yet.
English
Eugene Katrukha 🇺🇦 retweetledi

Virtual I2K starts TOMORROW, Monday the 28th! 34 FREE sessions, 900+ registrants so far. Learn more about favorite #bioimageanalysistools or find your next fav! i2kconference.org/virtual
Please also help our community improve by filling out our survey - forum.image.sc/t/2024-image-a…

English

I have no words to express how this saddens me; I was very much looking forward to being with you. I am hopeful I will be back on my feet tomorrow if I can. In my absence, please keep an eye on everyone from the lab:
@gomez_mariscal, @Bruno_MSaraiva, @IvanHCenalmor. They rock!
English

Hello #I2K2024 friends. You may wonder where I am and why I couldn't give a talk this morning. Unfortunately, I started feeling sick with nausea last night, and although I tried to push through this morning while still coming into HT, that ended up being a terrible idea (1/3)
English

@lhinderling I laughed, since that happened to me a couple of times. I hope you are not down, the main point is that you hopefully will arrive ok. And now you can show a presentation with movies instead of a boring 2D sketch.☺️
English
Eugene Katrukha 🇺🇦 retweetledi

How diverse is protein localization to human primary cilia? Check our study revealing incredible cell-type and single-cell diversity:
biorxiv.org/content/10.110…
3 year effort of studying 1935 proteins with imaging - 654 ciliary proteins pictured!
Tweetorial by @Prof_Lundberg & ⬇️
Emma Lundberg@Prof_Lundberg
I’m very excited to finally present something we’ve been working on for the past couple of years – a spatial proteome map of primary cilia, released in @ProteinAtlas today. Amazing teamwork led by @jn_hansen Check out our preprint doi.org/10.1101/2024.1…
English
Eugene Katrukha 🇺🇦 retweetledi

Announcing CT Foundation, a new medical imaging embedding tool that accepts a computed tomography (CT) volume as input and returns a small, information-rich numerical embedding that can be used to rapidly train models. Learn more and try it out yourself → goo.gle/4dYkClf
English
Eugene Katrukha 🇺🇦 retweetledi

Microscopy Nodes 2.0 is out! This includes a new channel interface and OME-Zarr support! 🔬🎉
youtube.com/watch?v=ybQcKE…
YouTube
English
Eugene Katrukha 🇺🇦 retweetledi

🚀 Thrilled to announce #inTRACKtive: a web-based tool for exploring massive cell-tracking datasets, no software installation required! Just open your browser and dive into terabytes of developmental biology data. We used it to build the virtual embryozoo.org of tracked embryonic development datasets 🐭🪰🪱🪲🐠, but you can also use it for your own data! 🔬
Preprint:
biorxiv.org/content/10.110…
Repository:
github.com/royerlab/inTRA…
This project was spear-headed by @TeunHuijben, together with engineers Andrew Sweet and @aganders3 from @cziscience
@czbbiohub #CZBiohubSF #devbio #tracking #web #visualization
🧵1/n
English
Eugene Katrukha 🇺🇦 retweetledi

2/ 🐠 Zebrahub-Multiome offers interactive data exploration via CZ #CellxGene and interactive figures powered by #Streamlit. Dive into the time-resolved and cell-type-specific complexities of chromatin accessibility and gene regulatory networks during zebrafish embryogenesis with unprecedented detail!
zebrahub.sf.czbiohub.org/epigenomics
#zebrahub #grn #devbio
English

Very pretty! And looks like just right dataset for #BigTrace analysis and visualization
English

@christlet @bio_physics No. Good addition, thank you. I saw the paper, but I was not convinced by the results shown in figures.
English

Hey tweeps ! Any favorite tool to segment filaments from images ? Better if it can handle crossing. Even better if it has tracking, but not a must.
(pref. in Fidji/Napari/Python, but Matlab is, I see ya old folks) @_malberto @katpyxa
English

@bio_physics Ok, here is the last one :) Similar idea to Fourier space decomposition, only they use pre-trained networks to separate orientations and link back github.com/i-yliu/mtquant
English

@katpyxa Thanks a lot !!! Will look at these options !
English

@bio_physics there is also this tool (matlab) that focuses on resolving intersections accurately github.com/mkitti/Adaptiv…
English

@bio_physics And there is also TSOAX/SOAX 2D/3D +T, but in C, standalone lehigh.edu/~div206/soax/i…
English

