KB Choi
639 posts


KB Choi
@kwangbom
Scientist in Computational Genomics, developing novel computational methods for high-throughput omics data. All views are my own. https://t.co/weMgLuJLvj



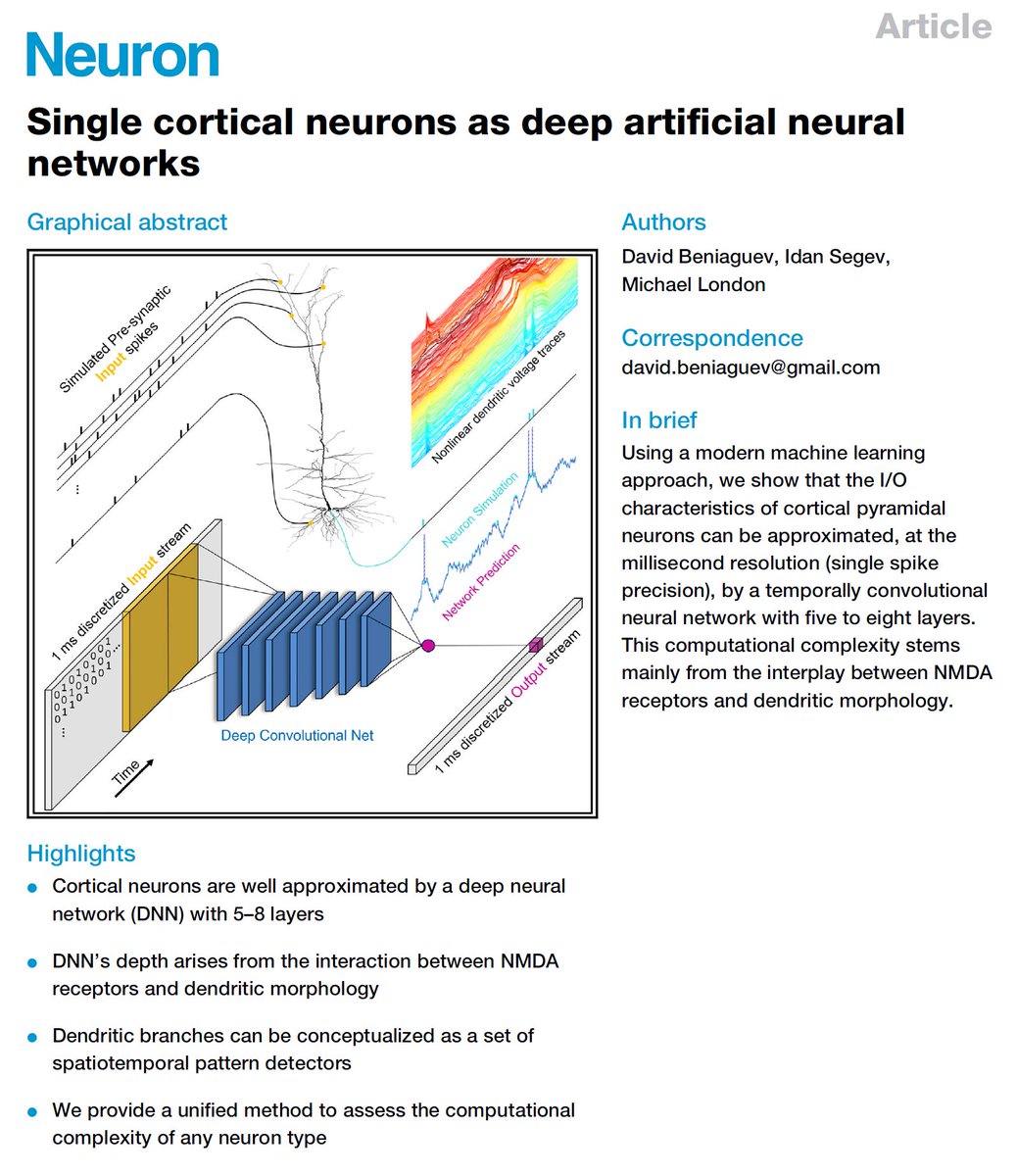
Unwritten rule for success in Academia: establishing your “brand” is very helpful on the job market. How do you do this?






Excited to share our lab's latest preprint, led by @CXchengxiangQIU, @bethkarenmartin & Ian Welsh of @jacksonlab. We set out to build a single cell roadmap for all of mouse prenatal development, from single cell zygote to free-living pup. Preprint: tinyurl.com/2nhe4mm9 1/n

Inferring super-resolution tissue architecture by integrating spatial transcriptomics with histology go.nature.com/47m0Mgk


Last day at Computer Science Building. I never realized how much one can get attached to the workplace. All your Ph.D. career you think when are you gonna graduate but when it's time you don't want to. Hopefully the plan to balance the emotions with Christmas is gonna work .

Ever analyze a scRNAseq dataset and wonder if a specific cell state has been seen before? And if so, where in the human body? Under what conditions? Well, now you can use our lightning fast SCimilarity search and foundational model for that! ⚡️🔎🧬 biorxiv.org/content/10.110… (1/11)

SIMBA learns a co-embedding space of single cells and multiple features such as genes, chromatin accessible regions, and transcription factor binding sequences, boosting performance of various analyses of cellular diversity and regulation. nature.com/articles/s4159…





Meet PyDESeq2 – a Python-based software package for #RNAseq differential expression analysis based on DESeq2. Owkin is open-sourcing PyDESeq2 to support the growing community of bioinformaticians adopting Python for their data analysis workflows. Check out PyDESeq2 on GitHub at github.com/owkin/PyDESeq2 Read more at owkin.com/en/publication… #bioinformatics #python


