
Matt Pahl
307 posts













GIP results are out. The beauty of human diversity in India hasn't faded a bit. Out of the 5000-odd communities, the largest genetic dataset for India includes only about 80 pops. This means these results are just 1.6% of the whole story 1/n #india medrxiv.org/content/10.648…

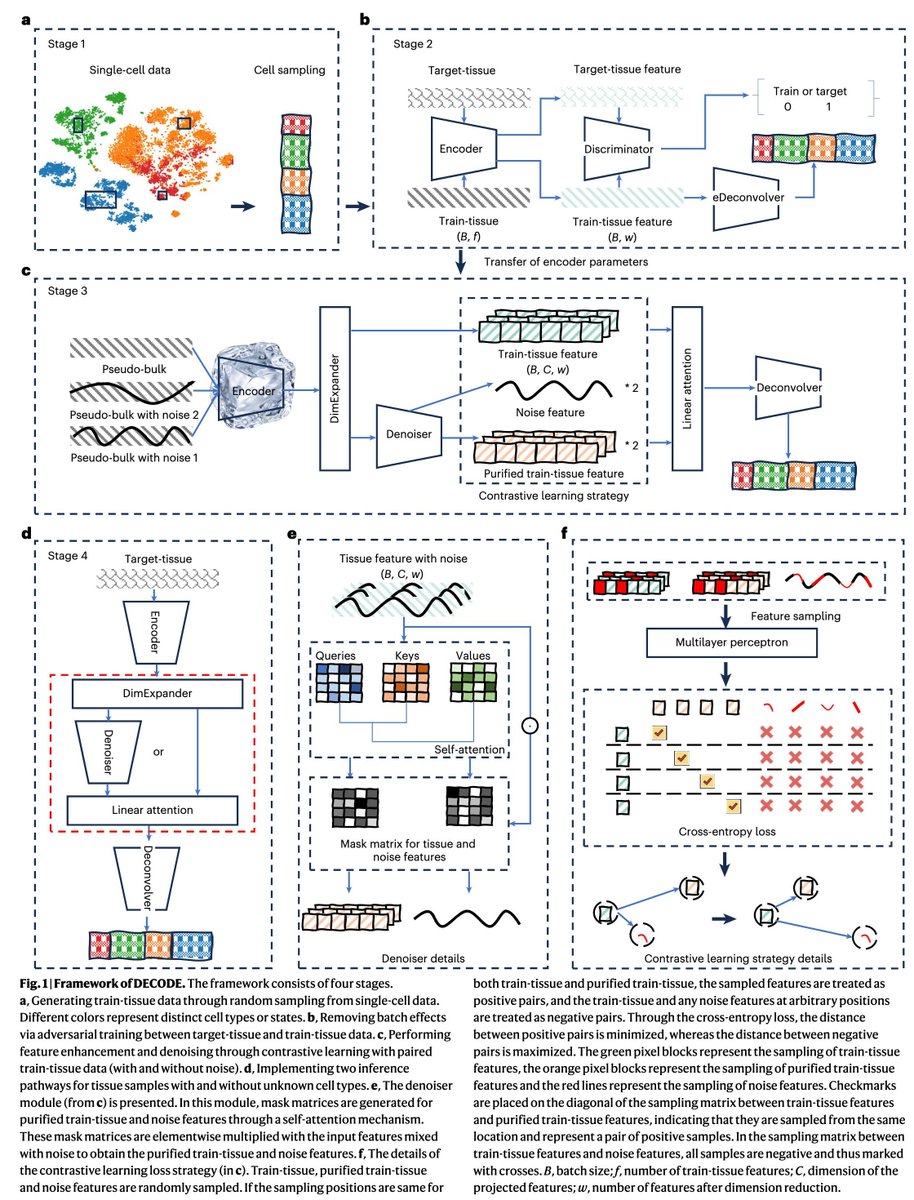
Thrilled to announce alphagenome-pytorch, an accurate, readable, and careful port of AlphaGenome's architecture and weights to PyTorch. Work with @gtcaa @m_kjellberg @chriswzou @tuxinming as part of the GenomicsxAI initiative between @anshulkundaje and @pkoo562 labs.







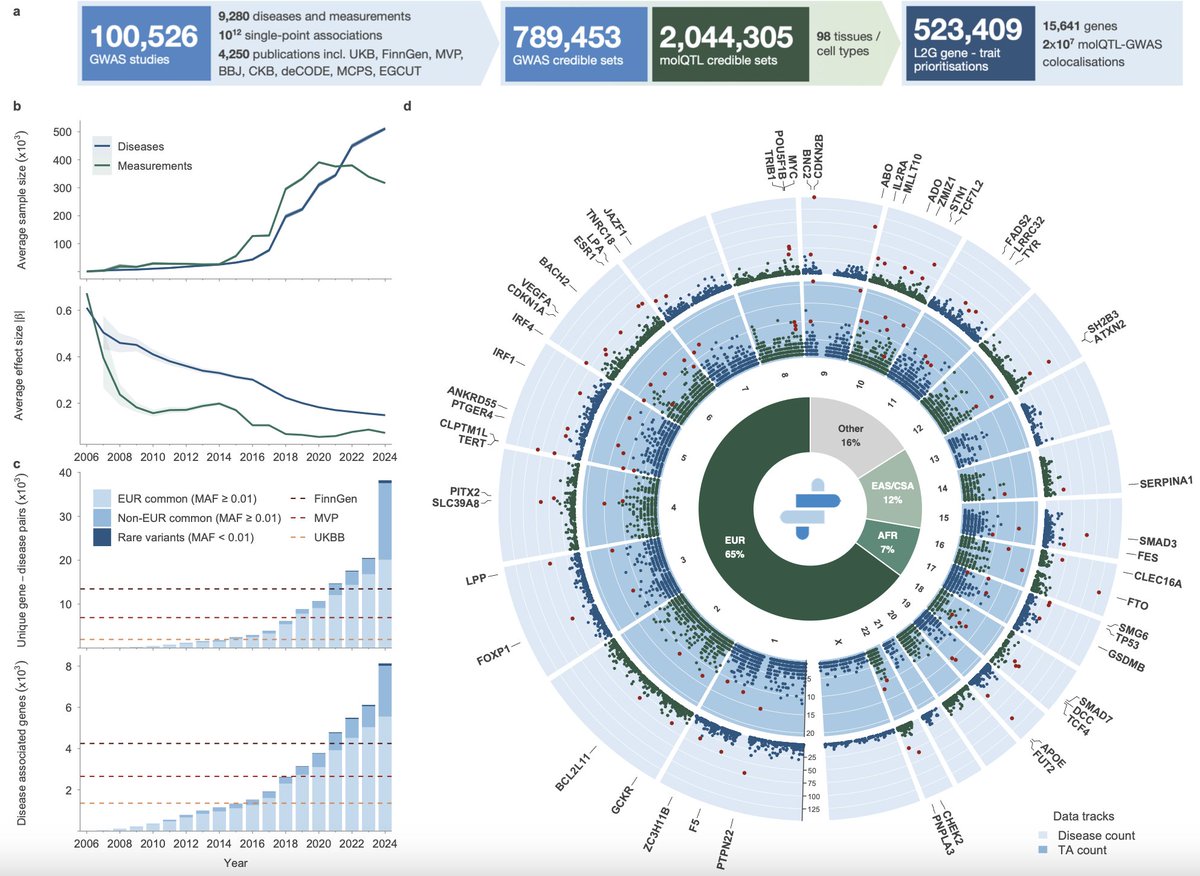
How do GWAS and rare variant burden tests rank gene signals? In new work @Nature with Jeff Spence, @jkpritch, and our wonderful coauthors we find the key factors are what we call Specificity, Length, and Luck! 🧬🧪🧵 nature.com/articles/s4158…

