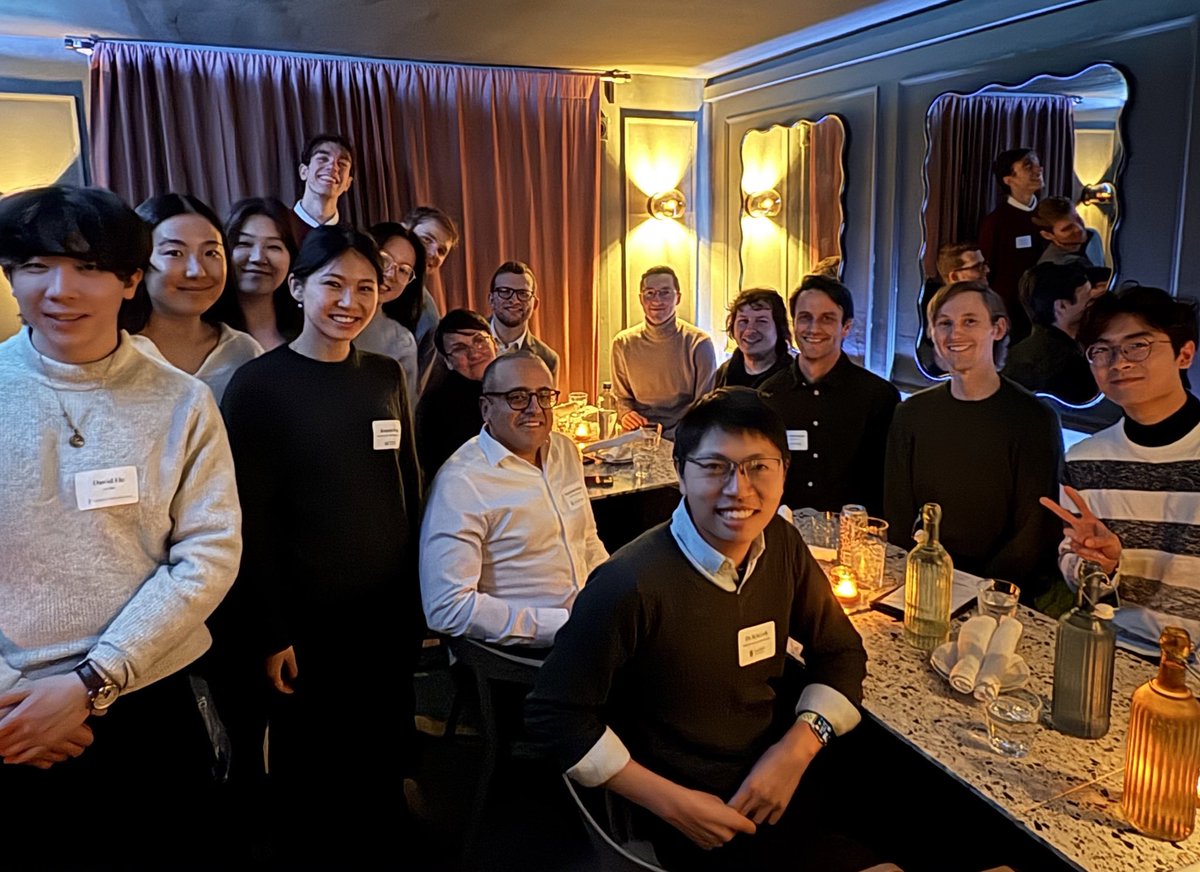
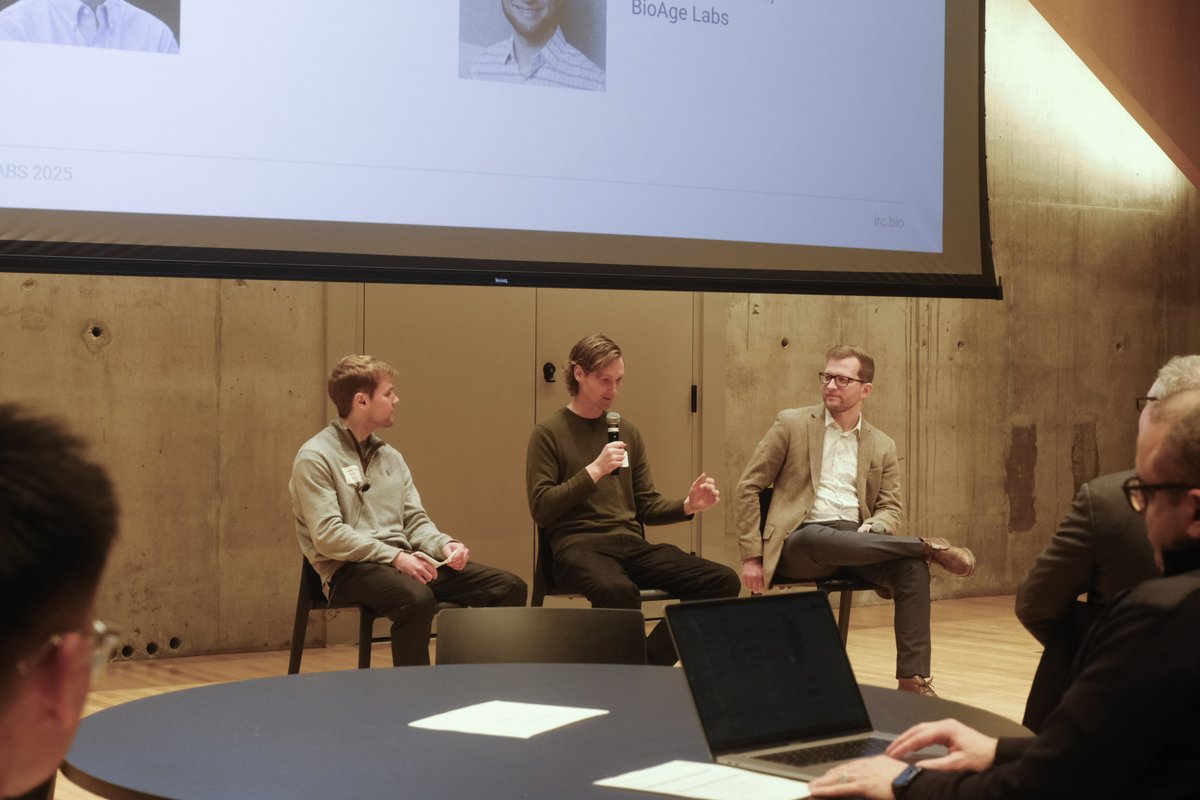
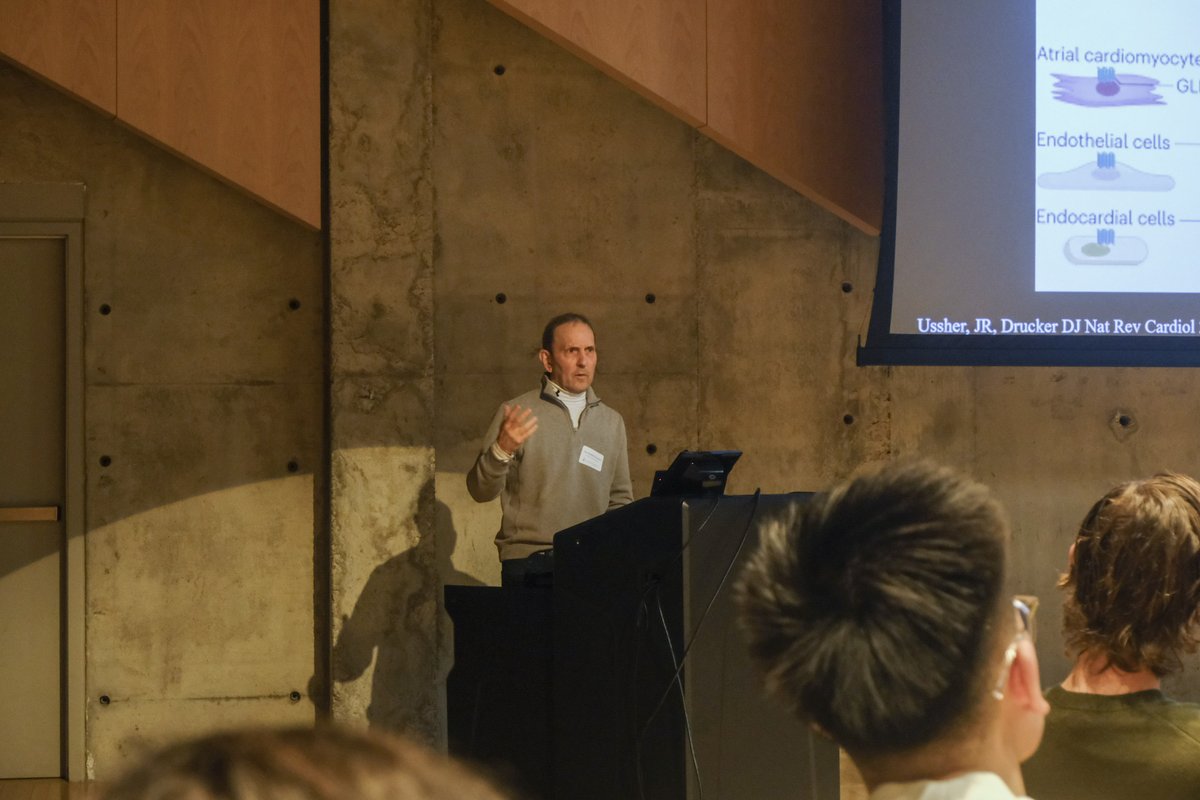
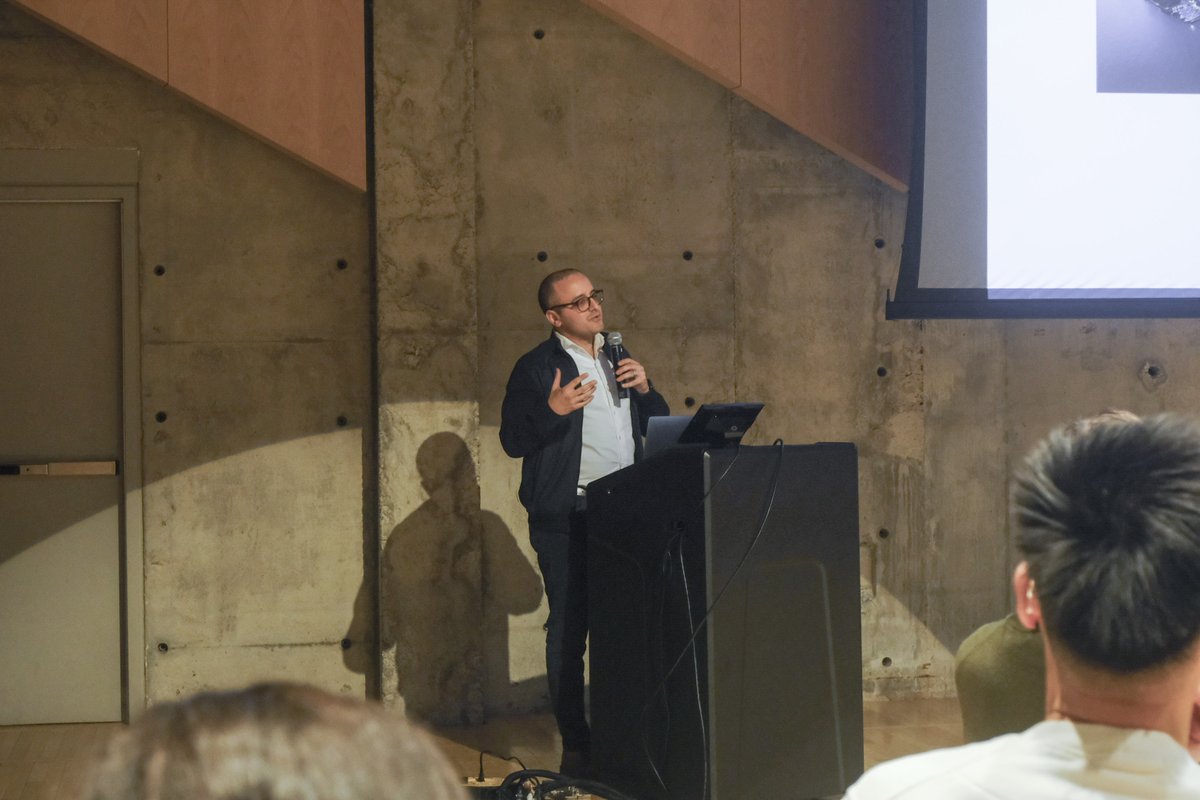
Jan 28, 2025 marked the 2nd edition of our annual Toronto Aging Biology Symposium. We are so grateful to have had 2 successful runs with nearly 300 registrations (undergrad, grad, alum, industry) this year. Furthermore we are ever thankful to our extraordinary lineup of speakers