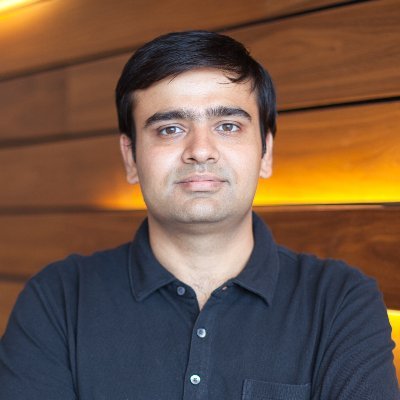

We mapped the zinc‑finger (ZF) “degrome.” High‑throughput ZF reporters + glutarimide analogs → new CRBN‑recruited degrons and design rules for degraders.
Shourya S Roy Burman
46 posts

@ssroyburman
Designing biological sequences | ex-@DanaFarber @harvardmed @JohnsHopkins @IITKanpur

We mapped the zinc‑finger (ZF) “degrome.” High‑throughput ZF reporters + glutarimide analogs → new CRBN‑recruited degrons and design rules for degraders.


















Nice DFCI Insight blog post about Sidney Farber Fellow @FranziskaWacht1 who will be conducting her research in @fischerlab1 blog.dana-farber.org/insight/2024/0…

Excited to share our story on polymerization in ZBTB transcription factors, which was published in the July 11 issue of @MolecularCell! cell.com/molecular-cell… 1/n





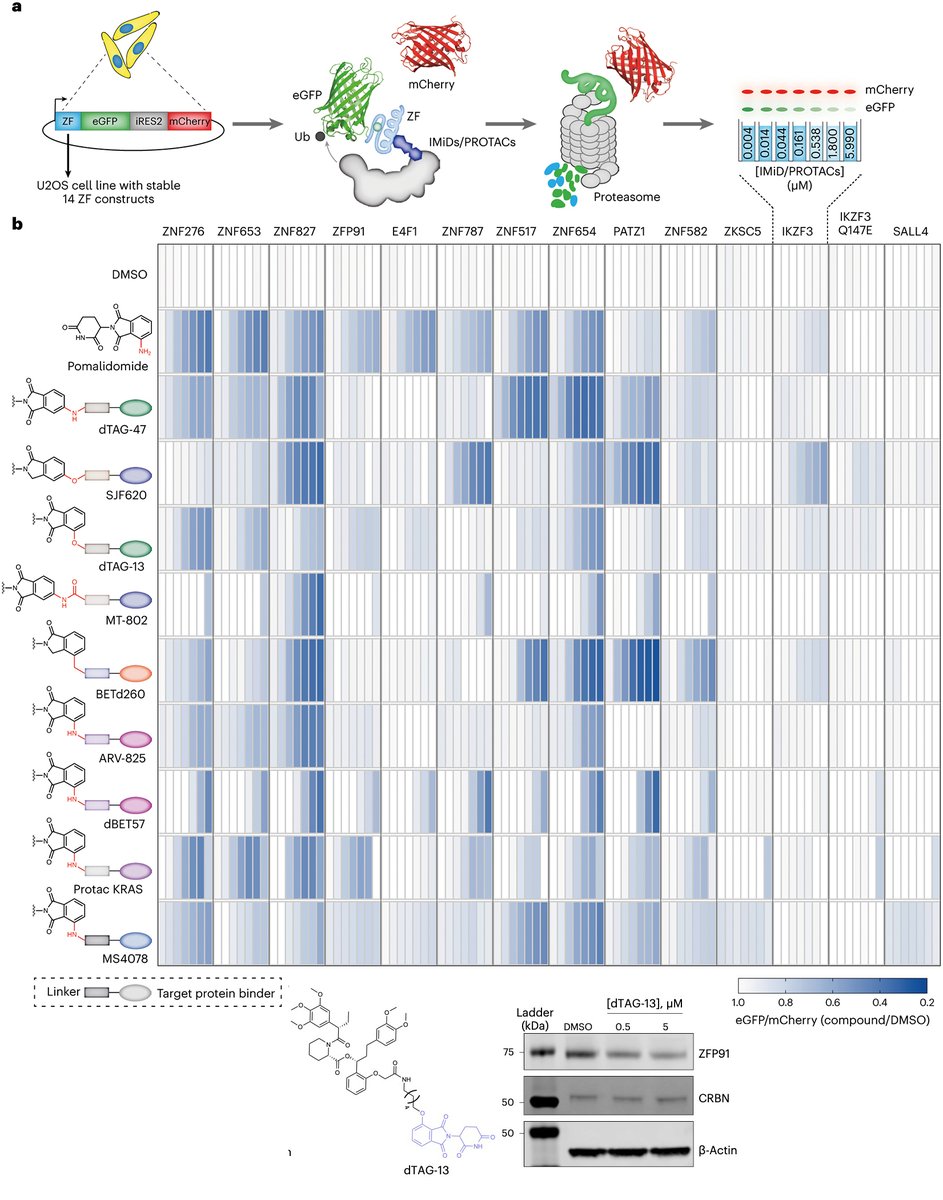

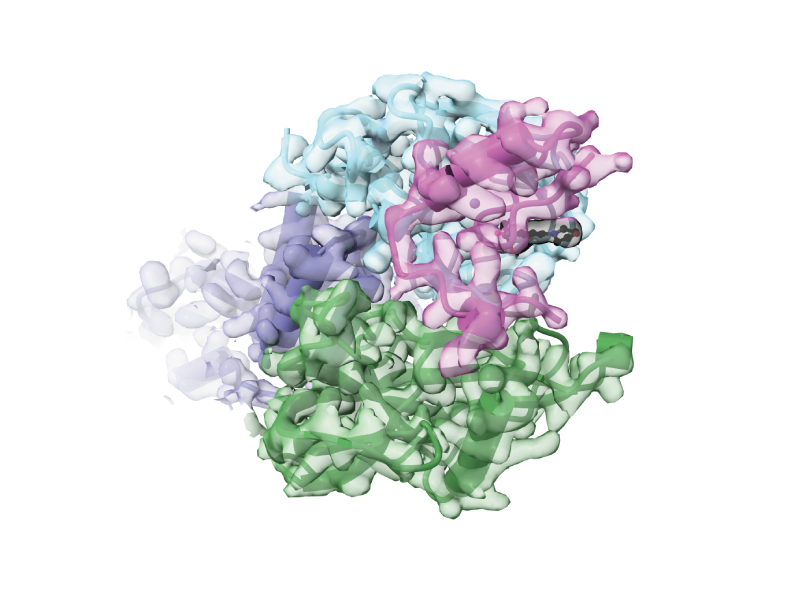



Today in @Science_Magazine we report the laboratory evolution of compact degrons that enable targeted protein degradation triggered by an otherwise-inert small molecule, a multidisciplinary study that integrates organic chemistry, molecular glues, protein evolution, genome editing, and structural biology. PDF: drive.google.com/file/d/18hu15h… 1/13