Tomoko Nakanishi
186 posts


@tomoco_naka
Respirologist, MD, and Ph.D student in Human Genetics




Nature research paper: The variation and evolution of complete human centromeres go.nature.com/3VJjaOg


We’re excited to share that the RGC-ME Browser is live! The RGC-ME Browser offers allele frequencies, ancestry estimates, constraint metrics, and transcript annotations based upon a catalog of human protein-coding variations from the exome data of ~1M individuals.

Genome-wide Association Study of Long COVID medrxiv.org/cgi/content/sh… #medRxiv







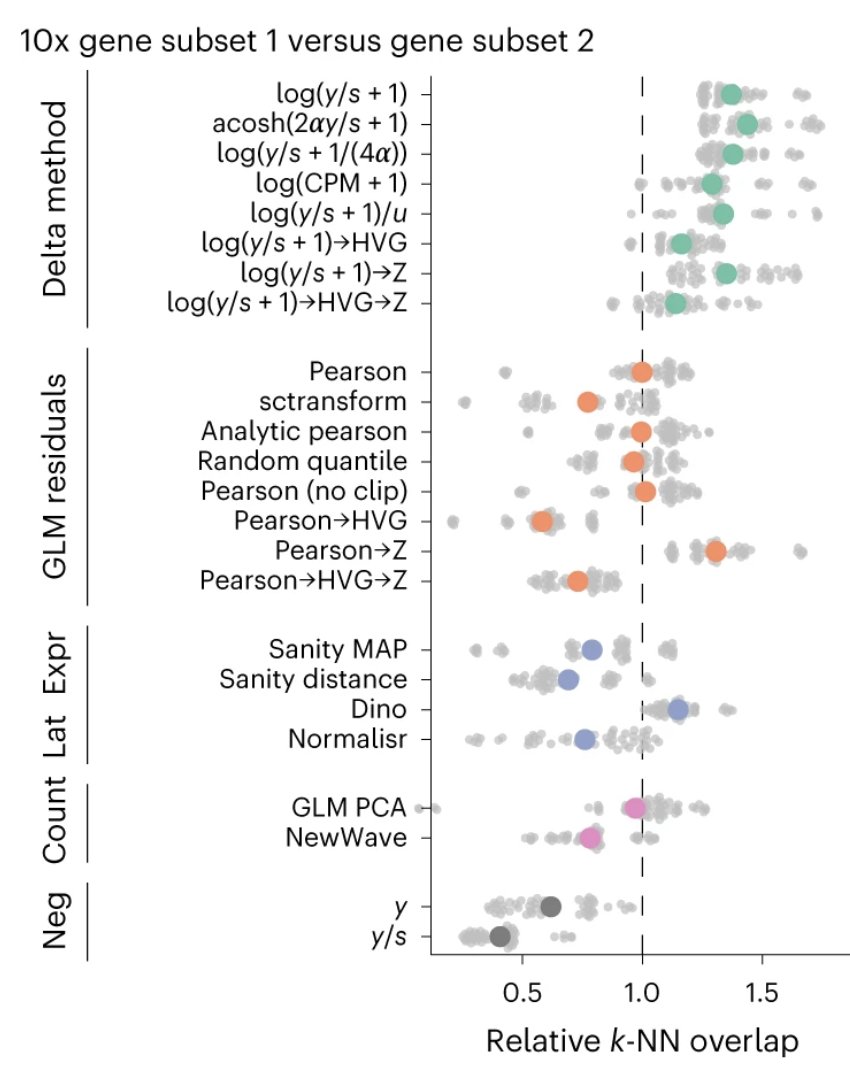


Sharing this in case it's useful: I find that students often have a hard time conceptualizing quantitative parameters in genomics and popgen Here's a cheat-sheet I put together for teaching. It's also part of a textbook I have been writing on human genetics (more on that below)