Yew Mun
775 posts

Yew Mun
@yew_mun
Father | Computational Chemistry Expert @ The Francis Crick Institute | Harnesser Opinions/Views are my own.



Introducing Orchids, the world's first vibe coding IDE. Orchids can build, watch, and listen on par with a human developer. Orchids ranks #1 on App Bench, the most rigorous benchmark for end-to-end software development. An agent, IDE, built-in browser, Supabase, and Stripe all in a single tool. Local, no lock in, no browser limitations - the next step forward in building with AI. Comment for 100k free credits at orchids [dot] app.
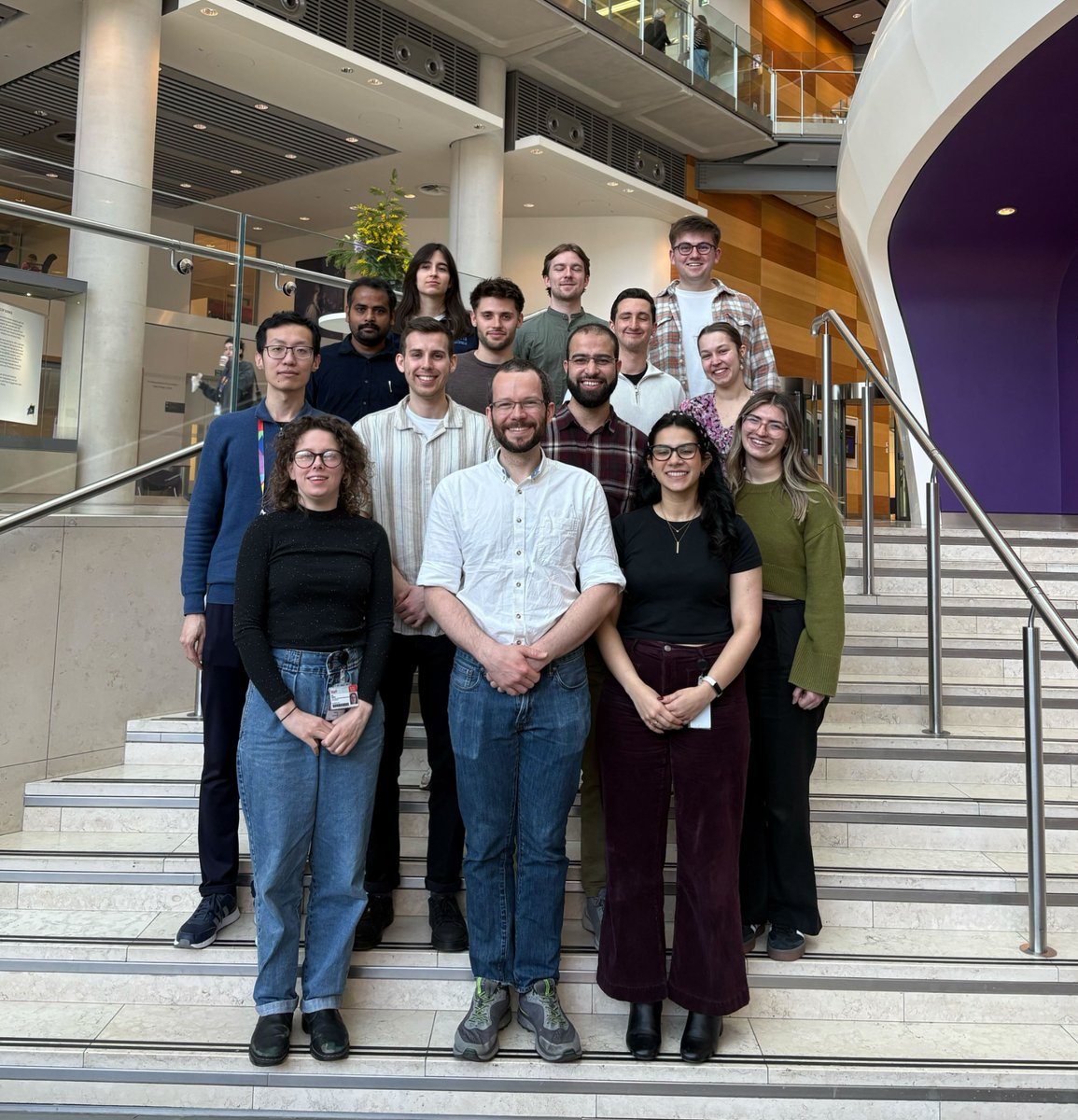



















Applications for @KingsCollegeLon @The_MRC Doctoral Training Partnership in Biomedical Sciences 2025 entry are now OPEN! Visit our website kcl-mrcdtp.com for more information. #MRCDTP2025




