

Kat
51 posts


@katyenko
building @bioArena_ AI ∩ bio | in vitro to in silico, always an experimenter



A few years ago, designing an antibody on the computer was extremely difficult. Today, there are several open-source tools which allow anyone to design antibodies from home. Out today: A step-by-step guide to antibody design. By @btnaughton.


Today we're launching free-energy perturbation (FEP) on Rowan! Using Rowan FEP, scientists can predict how tightly a potential drug will bind to a target, allowing teams to explore ligand modifications without having to wait for synthesis. Here's what this looks like:
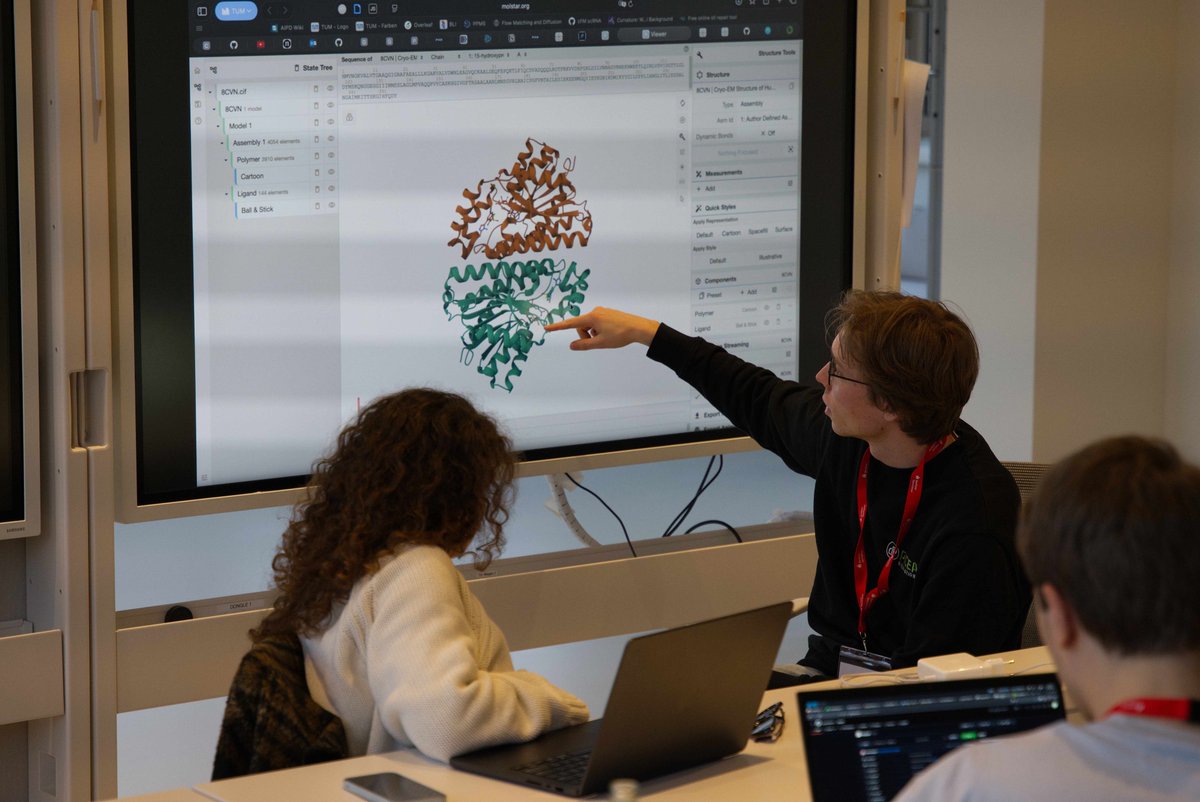
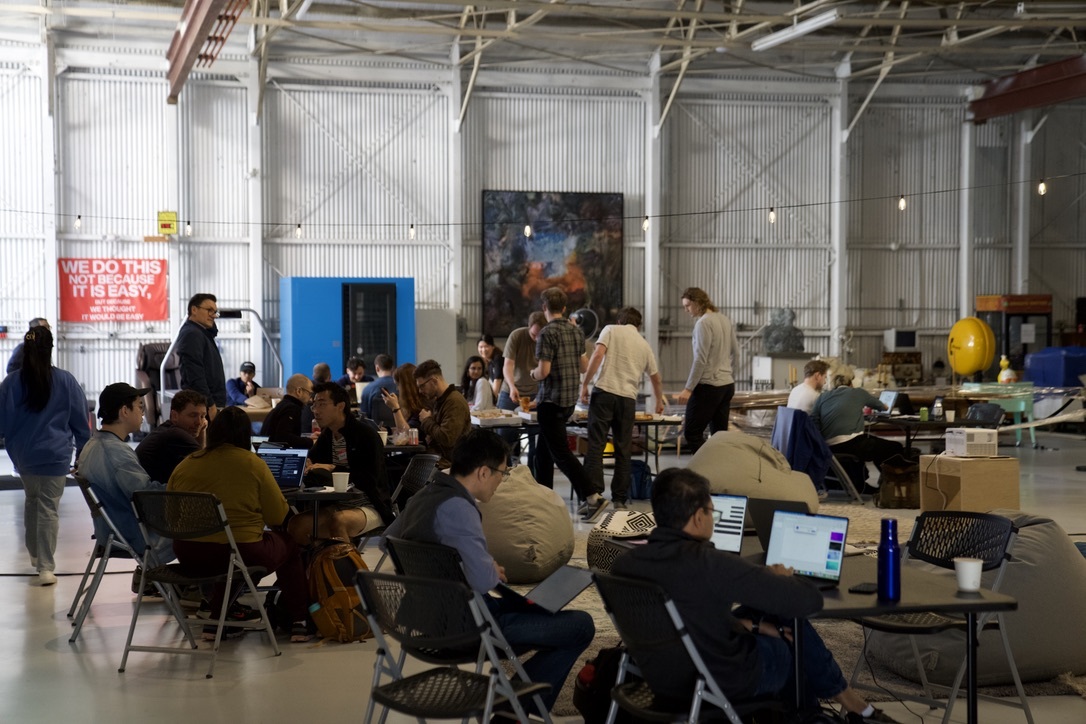








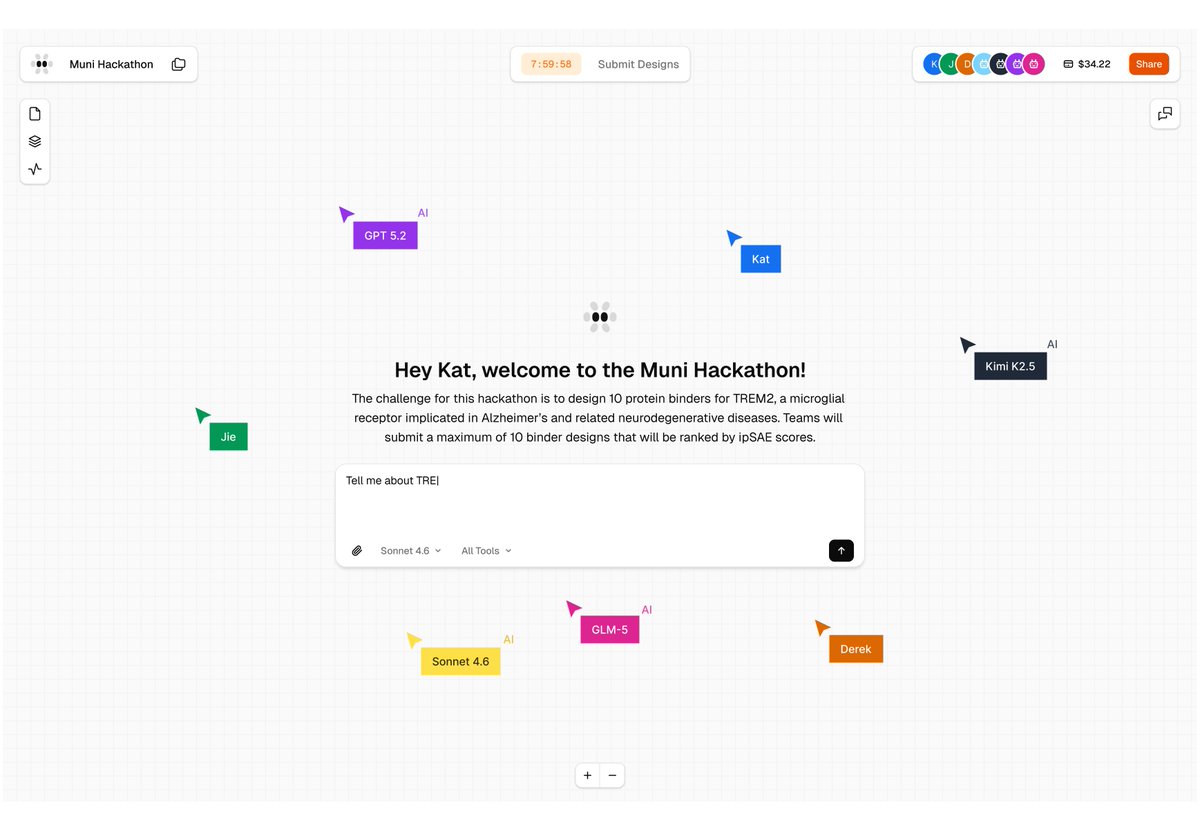

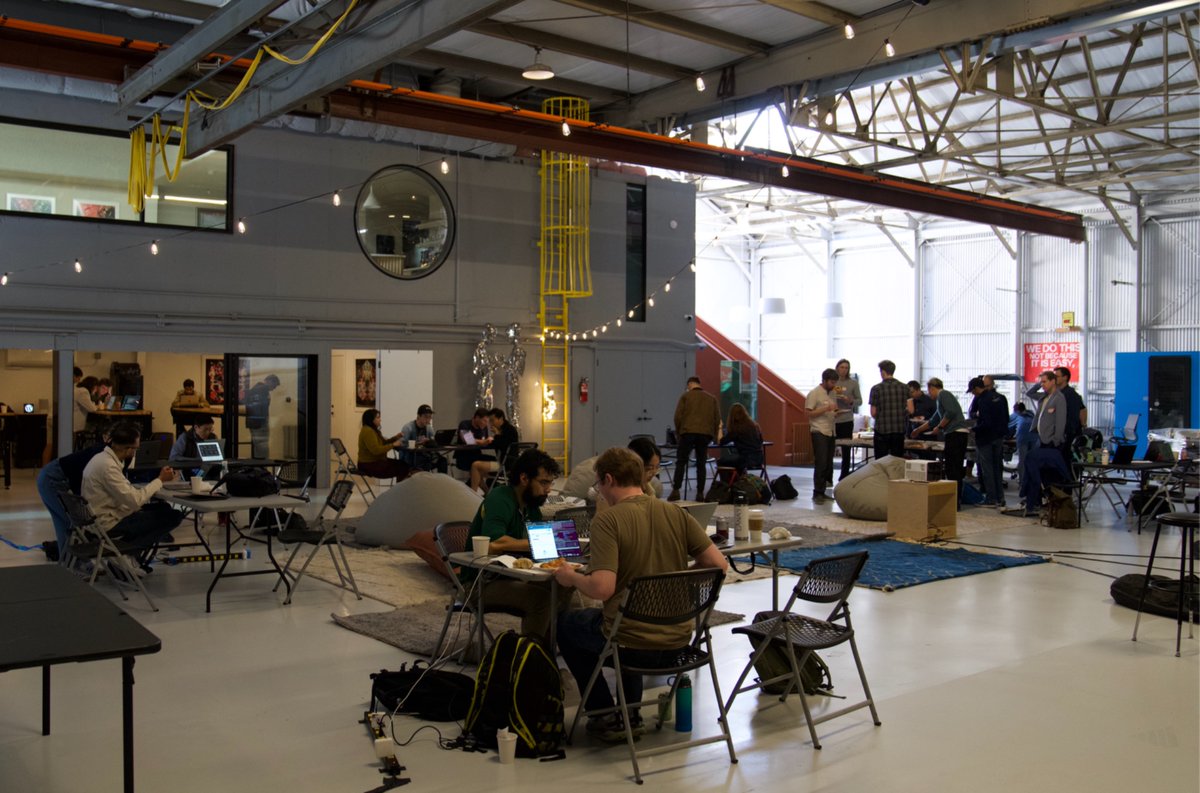
𝗖𝗮𝗻 𝗔𝗜 𝗮𝗴𝗲𝗻𝘁𝘀 𝗱𝗲𝘀𝗶𝗴𝗻 𝗽𝗿𝗼𝘁𝗲𝗶𝗻 𝗯𝗶𝗻𝗱𝗲𝗿𝘀 𝗼𝗻 𝘁𝗵𝗲𝗶𝗿 𝗼𝘄𝗻? 𝗛𝗼𝘄 𝗱𝗼 𝘁𝗵𝗲𝘆 𝗰𝗼𝗺𝗽𝗮𝗿𝗲 𝗮𝗴𝗮𝗶𝗻𝘀𝘁 𝗵𝘂𝗺𝗮𝗻𝘀? 𝗪𝗲'𝗿𝗲 𝗮𝗯𝗼𝘂𝘁 𝘁𝗼 𝗳𝗶𝗻𝗱 𝗼𝘂𝘁 🤖🧬 Teaming up with @bioArena_ for a single-day hackathon in San Francisco (Feb 28). The target: TREM2, a microglial receptor where loss-of-function variants increase Alzheimer's risk 2-4x. Alector's antibody AL002 failed Phase 2 last year. Novartis' VHB937 is still in trials. New binding modalities might find paths that existing antibodies missed. Human teams and autonomous agents compete side by side in this hackathon. Participants can submit to 10 sequences, ranked by the ipSAE score via Boltz-2. 🔬 Top 100 designs tested in our wet lab 📊 Results ~3 weeks post-event on @proteinbase 🤖 Get started with the Protein Design Skills for Claude Code → proteinbase.com/protein-design… Register → luma.com/a6t92ohv

𝗖𝗮𝗻 𝗔𝗜 𝗮𝗴𝗲𝗻𝘁𝘀 𝗱𝗲𝘀𝗶𝗴𝗻 𝗽𝗿𝗼𝘁𝗲𝗶𝗻 𝗯𝗶𝗻𝗱𝗲𝗿𝘀 𝗼𝗻 𝘁𝗵𝗲𝗶𝗿 𝗼𝘄𝗻? 𝗛𝗼𝘄 𝗱𝗼 𝘁𝗵𝗲𝘆 𝗰𝗼𝗺𝗽𝗮𝗿𝗲 𝗮𝗴𝗮𝗶𝗻𝘀𝘁 𝗵𝘂𝗺𝗮𝗻𝘀? 𝗪𝗲'𝗿𝗲 𝗮𝗯𝗼𝘂𝘁 𝘁𝗼 𝗳𝗶𝗻𝗱 𝗼𝘂𝘁 🤖🧬 Teaming up with @bioArena_ for a single-day hackathon in San Francisco (Feb 28). The target: TREM2, a microglial receptor where loss-of-function variants increase Alzheimer's risk 2-4x. Alector's antibody AL002 failed Phase 2 last year. Novartis' VHB937 is still in trials. New binding modalities might find paths that existing antibodies missed. Human teams and autonomous agents compete side by side in this hackathon. Participants can submit to 10 sequences, ranked by the ipSAE score via Boltz-2. 🔬 Top 100 designs tested in our wet lab 📊 Results ~3 weeks post-event on @proteinbase 🤖 Get started with the Protein Design Skills for Claude Code → proteinbase.com/protein-design… Register → luma.com/a6t92ohv

@NPraljak @IsomorphicLabs Well it is not 0 on our test and evaluation sets, I'm not sure what went wrong here...






In Silico to In Vitro is our most ambitious hackathon yet. We’re teaming up with @adaptyvbio to run a challenge where scientists generate novel binders in silico, with top designs built and tested in the lab. Join us if you're in SF on 2/28: luma.com/a6t92ohv Co-sponsored by @RowanSci, @openrouter and @modal, with generous compute support.
