固定されたツイート

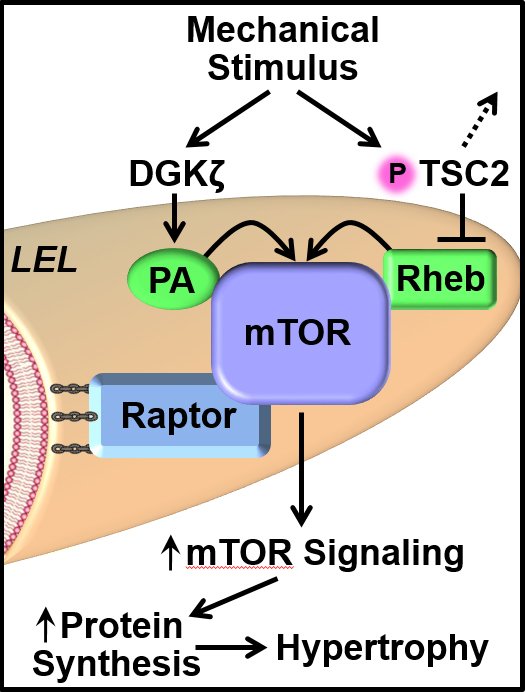
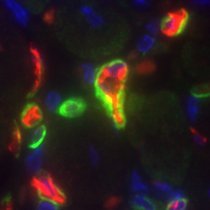
I'm very happy to have our #endogenous #SkM #EV paper out in @AJPCellPhys. journals.physiology.org/doi/epdf/10.11… A beautiful piece of art inspired by our paper (by Andrew Crowe) was selected for the cover. A 🧵 about the science (and our growing pains as a new lab in a new (to us) field)...

English