
𝓟𝓮𝓷𝓰 𝓗𝓮 何鵬
1.1K posts


@PengHeAtlas
bioinformatics for multiome➕spatial➕single-cell; development↔disease; Assistant professor at UCSF; Tweets👉own; BlueSky: https://t.co/5yOclEVU7u Mastodon:penghe











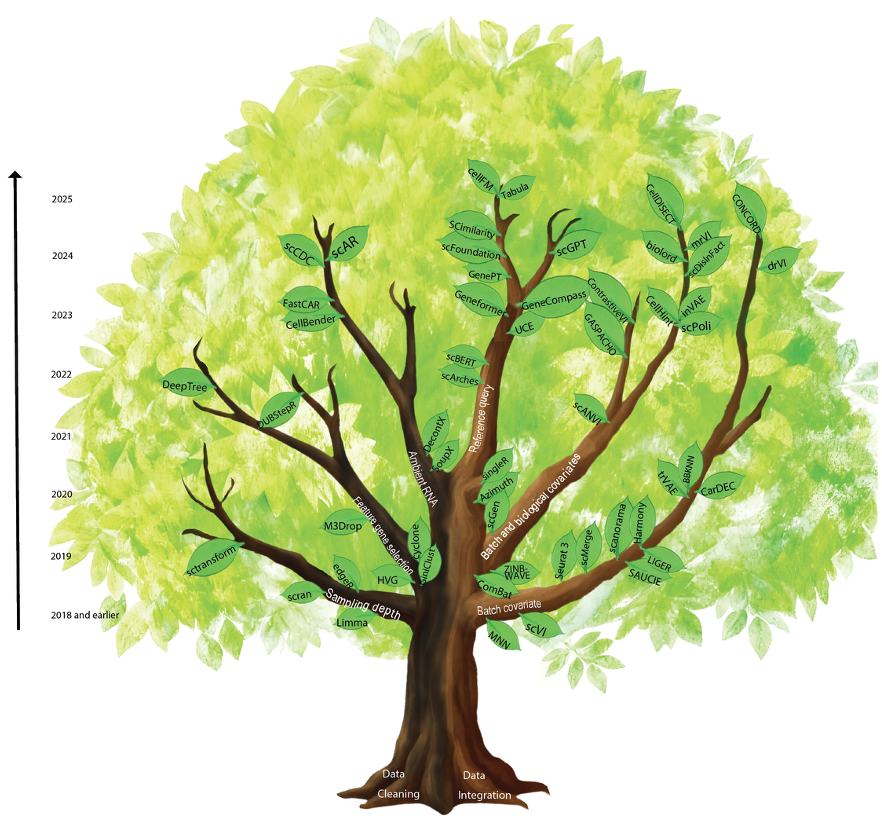
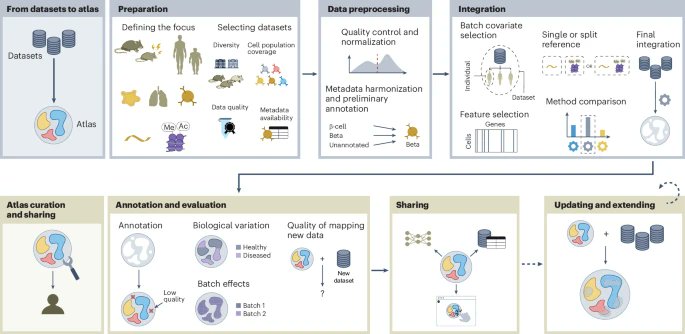
Defining “what is good” is central to both AI and Science. Despite massive efforts towards building the Virtual Cell, our recent @NatureBiotech article reflects on whether current gold standards (i.e., evaluation metrics) still hold up. nature.com/articles/s4158…




We are hiring! Join us in building virtual embryo models — an ambitious and exciting mission that goes beyond the widely acclaimed “virtual cell” initiative! 🚨 Are you a technology developer, developmental biologist, or machine learning expert interested in creating predictive, virtual 3D mammalian embryos to advance human health, particularly for congenital heart diseases? We’re looking for applicants with expertise in genomics, developmental biology, generative AI, or related fields to join our visionary project. The selected candidate will work within a multidisciplinary team of biologists, engineers, machine learning experts, mathematicians, and physicists. We offer an enriching, collaborative environment where post-docs, students, and team members can develop into leaders in both academia and industry. Join us on this exciting journey to shape the future of human health! ! 🚀 Nature career link below: nature.com/naturecareers/… Email: xiaojie at stanford dot edu